Diamond Workshop 2017
This page is provided as a generic orientation of the contents of the practical hands-on sessions in Diamond (November 2017).
Contents
Computers
Entering the terminal
The login and password are written in the front board.
Starting a Dynamo session
Dynamo is already preinstalled in the file system of Diamond. You can activate it through
module load EM/dynamo
As some visualizations are performed with Chimera you need to activate it as well
module load chimera
After Dynamo and Chimera have been loaded, you can start a session with:
dynamo
Bringing the data to the local machine
Some tutorials during the workshop require downloading big files from the Dynamo wiki using wget. You don't need to do that, as they have been already downloaded into the location /dls/tmp/DynamoWorkshop/data, which is visible by all the computers in the workshop.
To speed the access to these data sets, it is convenient to make a local copy of them into your own machines. To do so, defore going to lunch, please launch the order:
/dls/tmp/DynamoWorkshop/data/fetchDataSets.sh
from a linux shell. This will copy the data into the directory ~/dynamo/data in your machine.
Program
General Introduction
Guided presentation:
- tutorial on basic elements: help, data and metadata formats.
- tutorial on the basic concept in Dynamo alignment: the project.
Working on your own:
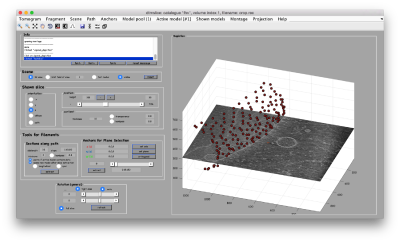
- Basic walkthrough: creating a catalogue, picking particles, launching a project.
- Advanced starters guide on a real tomogram. (~2 hours)
- Further work:
Geometric modeling
Short guided presentation:
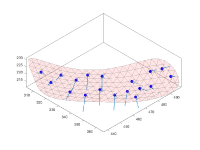
- tutorial on membrane modeling with dmslice
- Filament models with dtmslice
- Reusing model workflows ( walkthrough)
- Further work: catalogue
Working on your own:
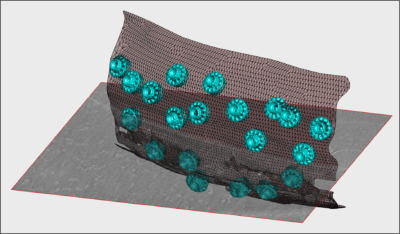
- In the afternoon, after the research talks, we will focus on the extraction of particles from densely packed spherical geometry on HIV viral capsides (~1 hour)
Template matching
Working on your own:
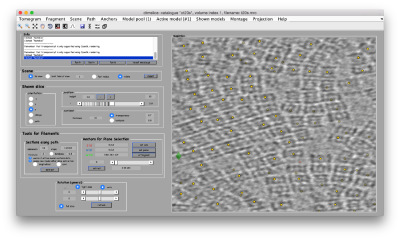
- We will follow this walkthrough for automated identification of proteosomes on a real tomogram through template matching. (~1 hour)
Adaptive bandpass filtering
Working on your own:
- We will follow this walkthrough to create a small synthetic data set that illustrates the principles of adaptive bandpass filtering, a way of conducting a golden standard alignment procedure . (~40 mins)
Classification
Short guided presentation:
- PCA Basic concepts.
- Multirefererence Analysis
Basic concepts.
walkthrough (~10 min) - Further work
- Commandline operations with PCA
Creation of 3D scenes
Working on your own:
- Walkthrough on the FHV data set (~1hour)
Further support material.
- Walkthrough on depiction and manipulation of triangulations (synthetic data).
Additional tools
Wednesday afternoon session.
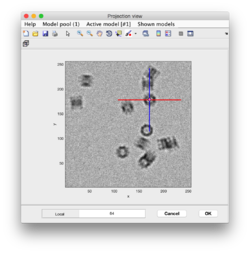
- Subboxing
- motivation slide: extraction of vertices from icosahedral viruses
- Basic subboxing (tutoriall)
- advanced subboxing tutorial: combining with MRA tutorial
- suggested data set: prd1 Capsides (in the folder of isolated particles)
- Manual alignment.
- Exercises
Data sets
The data sets will be updated here.
Exercises
Exercises are scheduled for the last day. Nevertheless, feel free to start them any time during the workshop.