Walkthrough on adaptive bandpass filtering
The adaptive bandpass filtering technique encompasses two elements:
- independent refinement of two half sets of the data
- dynamic computation of the low pass filter at each iteration
Contents
Creating a test data set
We will create a small data set of 50 particles to visualize the convergence process when using ABP. dtutorial can be used to produce random rotations and shifts of a given template, in this case a cube of 32x32x32 pixels containing a ribosome template
dtutorial ttest2 -M 50 -noise 0.2 -tight 1 -template ribosome32.em -dshift 0
- -dshift 0 ensures that the randomly rotated templates are not shifted
- -tight 1 forces the synthetic ribosomes to stay in a box of 32 pixels of sidelength
- -noise 0.2 adds gaussian noise of the given amplitude to each synthetic particle
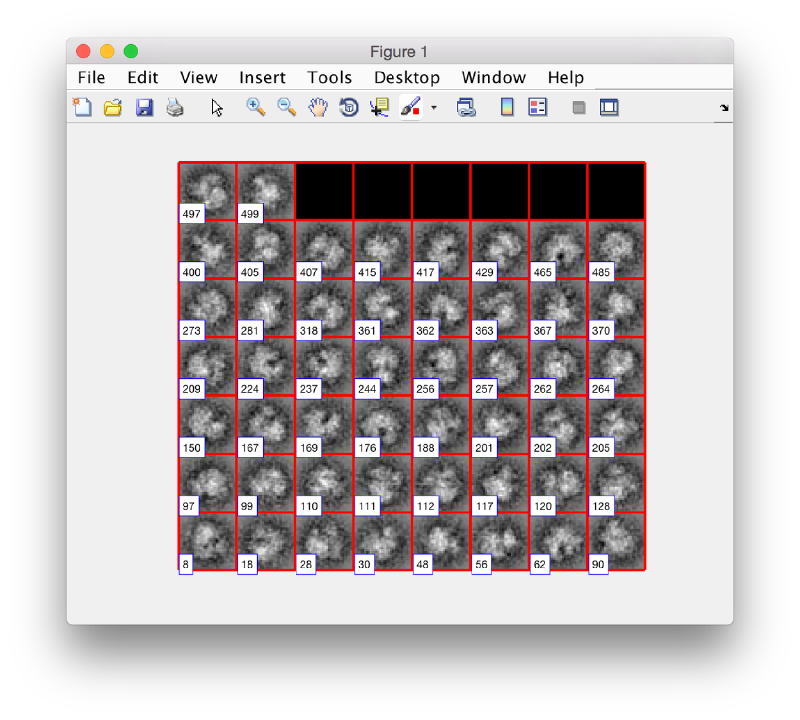
We can check the produced volumes:
dslices ttest2/data -jz 0 -labels 1
Creating an ABP project with the GUI
The easiest way to create an adaptive bandpass project is to first create an auxiliary single reference project and then deriving the corresponding ABP project.
Creation of supporting single reference project
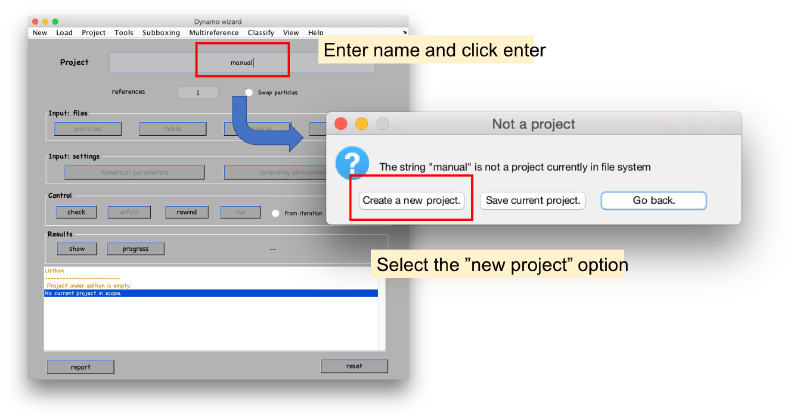
Launch the project editor through:
dcp;
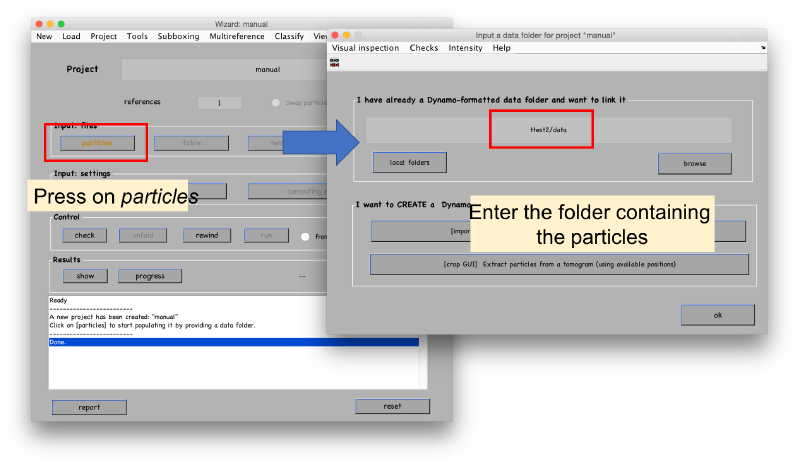
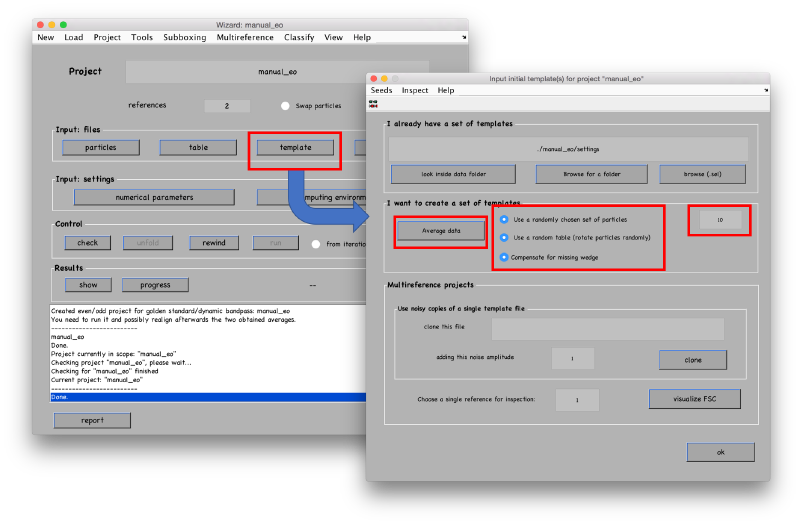
The different fields will open as we input the required info. We start with the particles.
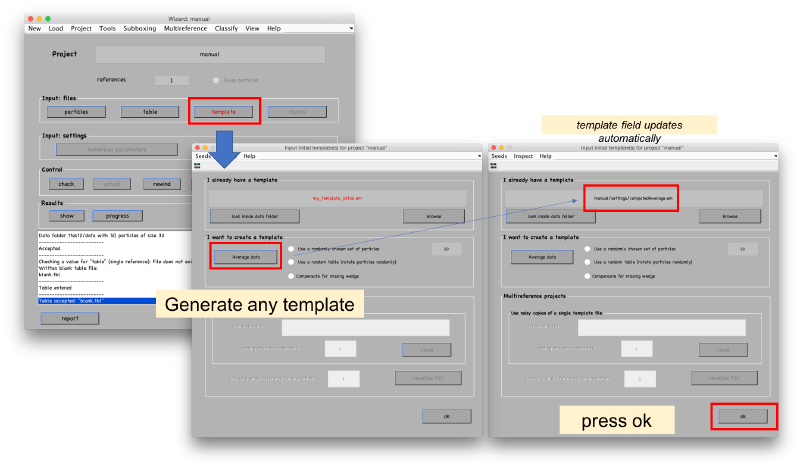
Seed files
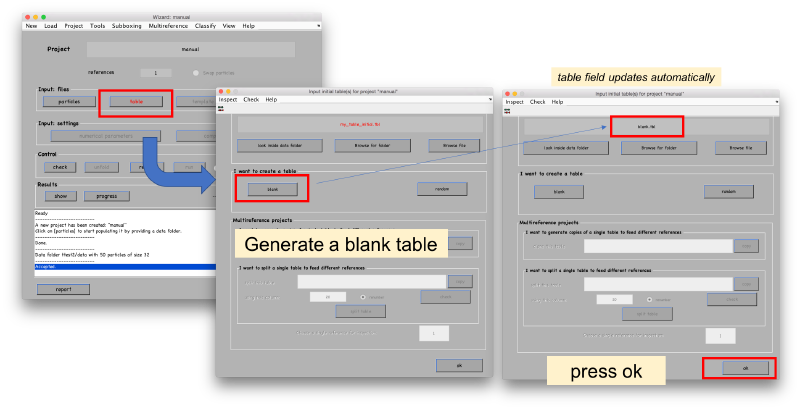
Table, template and masks don't need special formatting. We will just provide Dynamo with the default options. The table can be initialized to a blank table, which just contains the indices of the particles.
Template can be initialized just as an average of particles. This average will be recomputed anyway when we derive the ABP project.
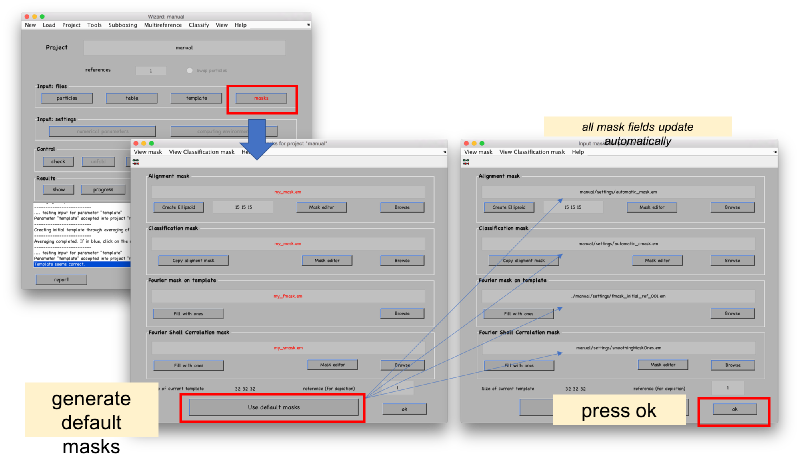
Masks are initiated to default values. This means spherical masks for alignment and classification masks, and full cubes for smoothing and fourier mask of template.
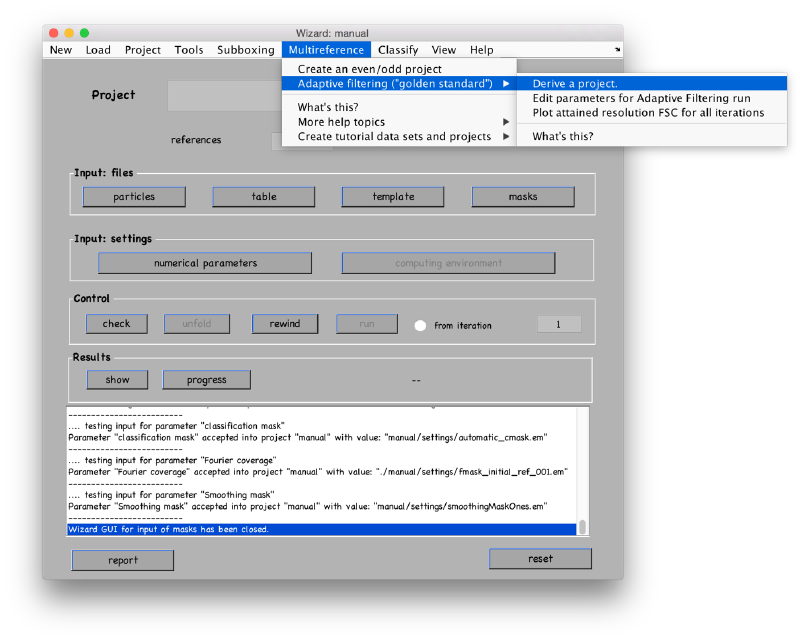
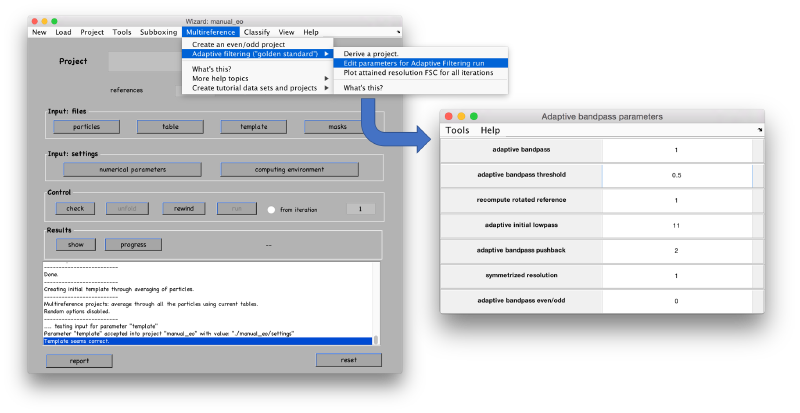
Derivation of ABP project
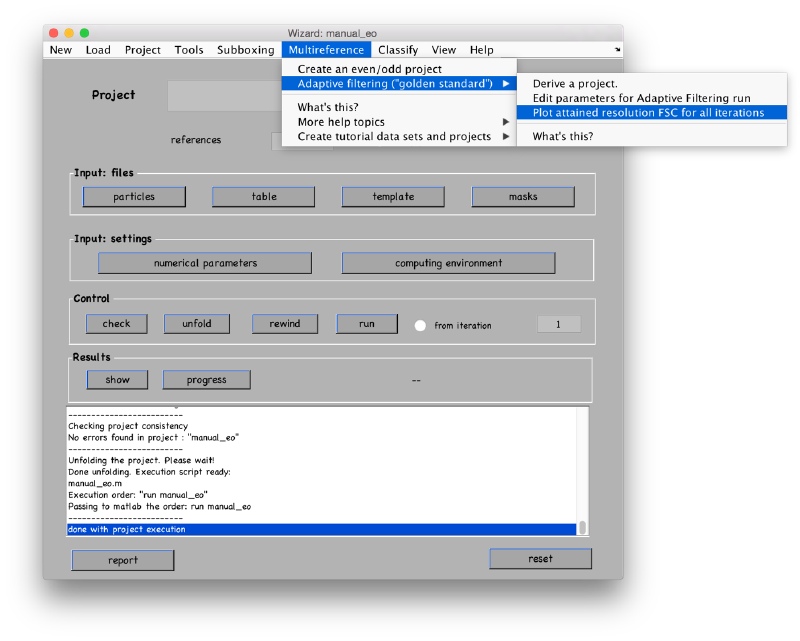
After the initialization of the support single reference project, you can derive an ABP project, through Multireference > Adaptive filtering > derive a project "
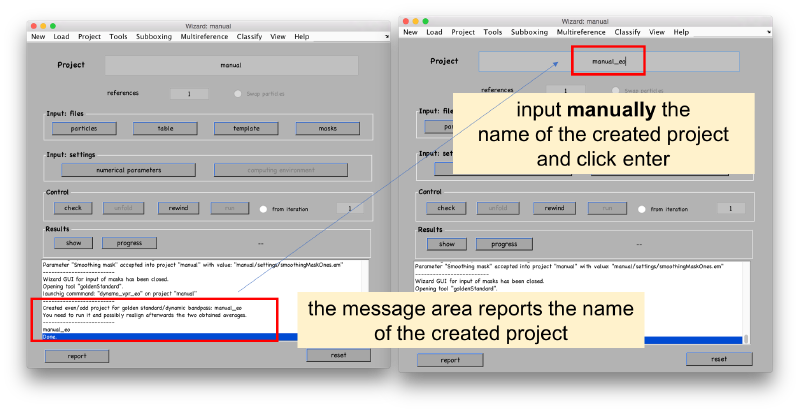
You need to edit the created new project (by default will have the same name as the base project with the string _eo appended)
In this new project, the table in each reference is indexing a different set of particles. The initial template will be computed independently for each of the two tables. As we are not going to use any a priori information coded in the tables, we ask Dynamo to use a random set of orientations. We also fix a small number of particles which will be randomly selected on each table to produce one template on each of the two channels.
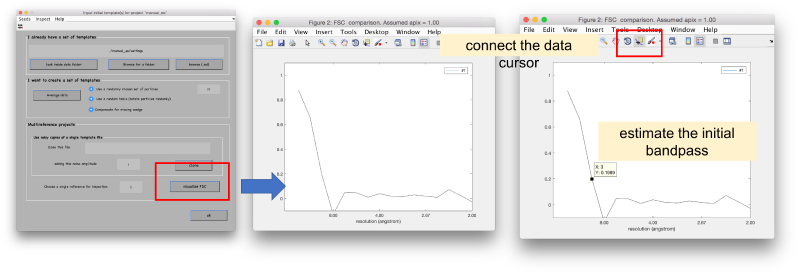
We can measure the similarity of the two initial templates in the Fourier space, and impose it later as starting value for the bandpass.
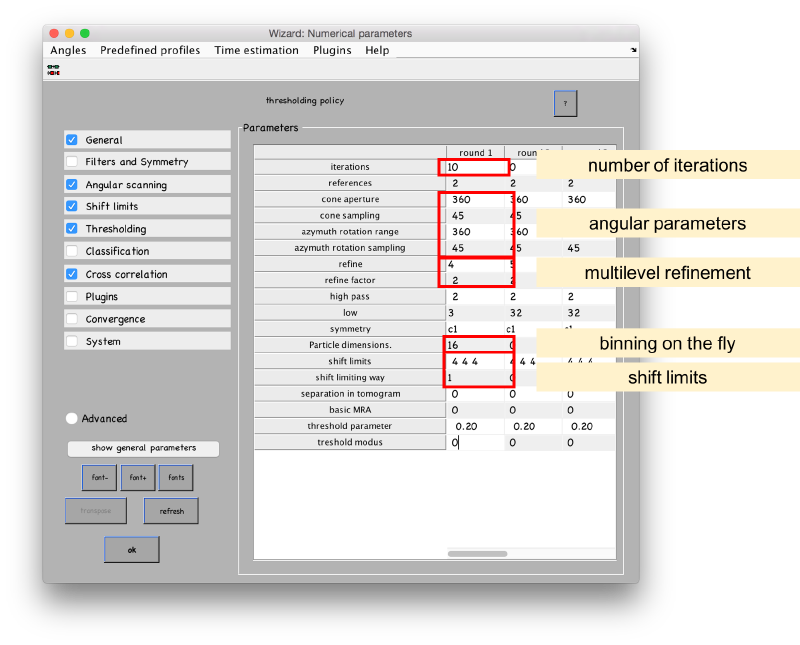
ABP-specific parameters can be found in the Advanced area of the Numerical parameters GUI. However, they are faster accessed through the Multireference > Adaptive Bandpass > Edit parameters " menu option.
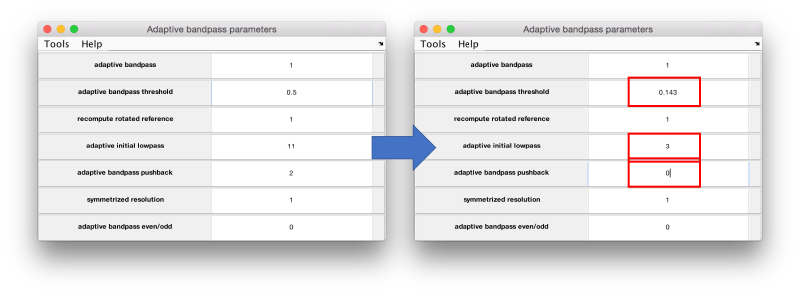
The main parameters in this window are
- initial bandpass. A safe option in view of the FSC is 3 pixels.
- threshold. Put it to 0.143 in this experiment.
- pushback. Should be set to 0. The default assumes a very conservative policy that can slow down the convergence process.
In the numerical parameters, we configure 10 iterations along these lines:
- We will use a rough angular sampling (steps of 45 degrees in both angular parameters) on the full rotational space (360 degrees in both angular parameters).
- A local angular refinement will be used on each angular scan. It will be used 4 times, each times halving the angular interval.
- We don't let the parameters drive away from the center of the particle (limit modus 1) more that 4 pixels.
- The original particles have a sidelength of 32 pixels. We impose a sidelength of 16 to bin them and accelerate the computations.
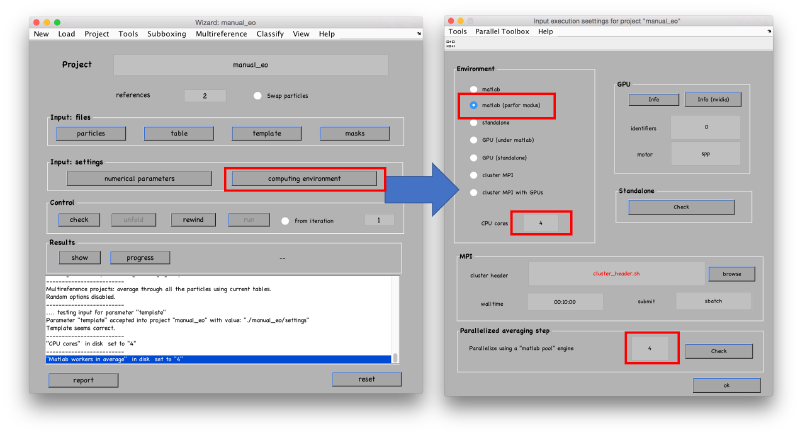
Then select the number of available cores for alignment and averaging.
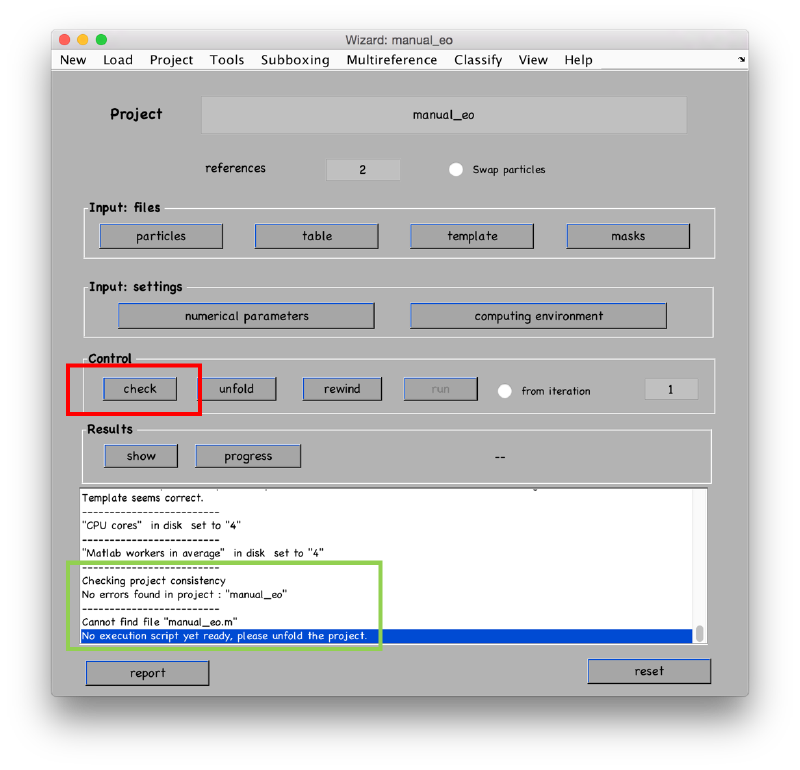
Check that the project can be executed:
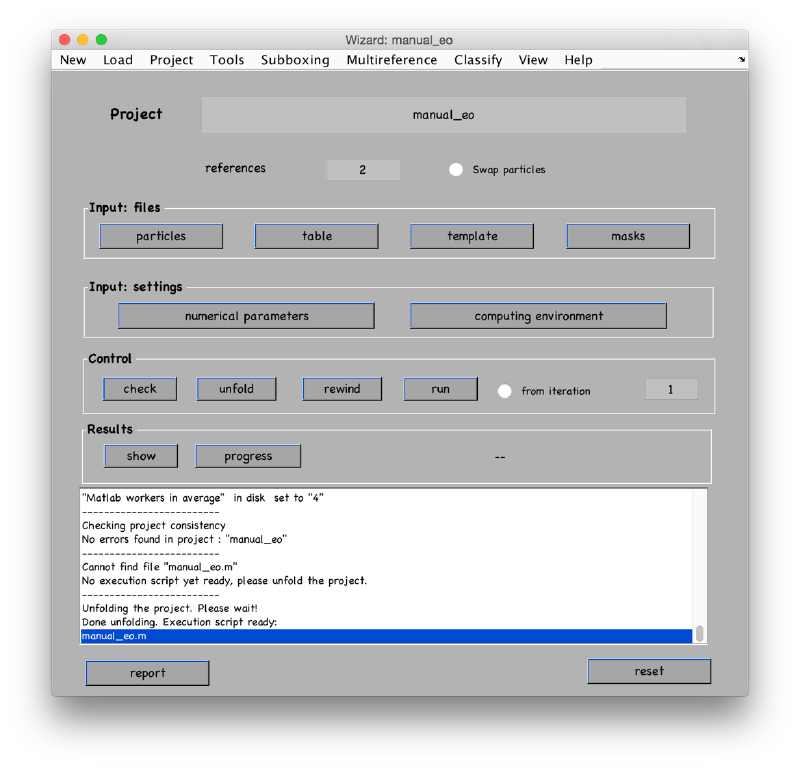
Prepare the project for execution:
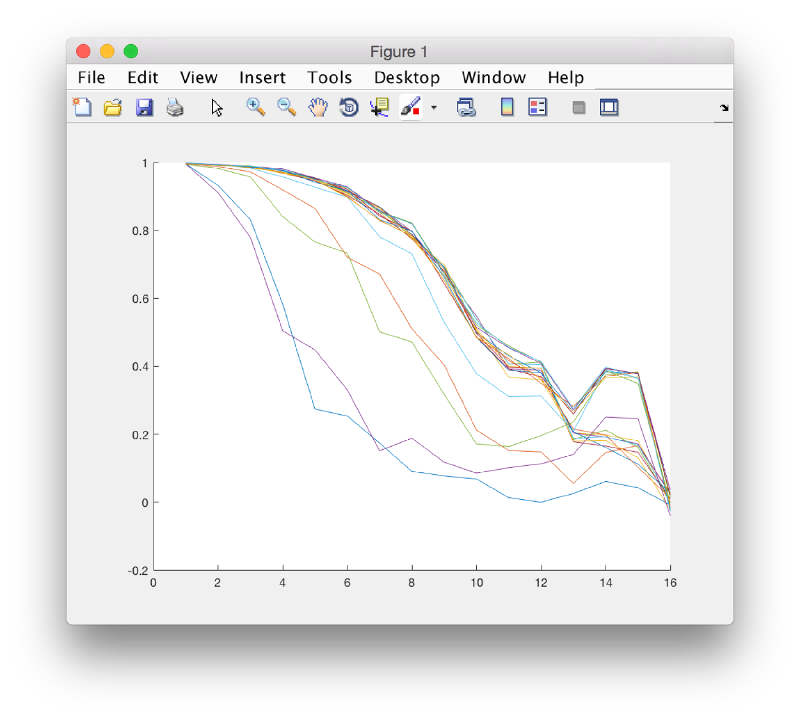
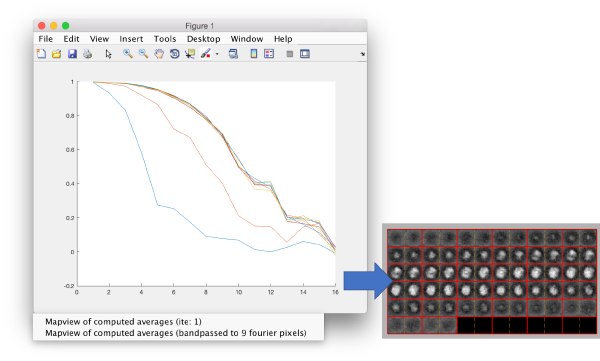
After completion of the project, we can check the FSC between the averages of the half data sets on each iteration.
Ignore the angstrom label: this is only valid when you enter the actual pixel size of the particles into the project. We observe how the last iterations don't move to higher resolutions.
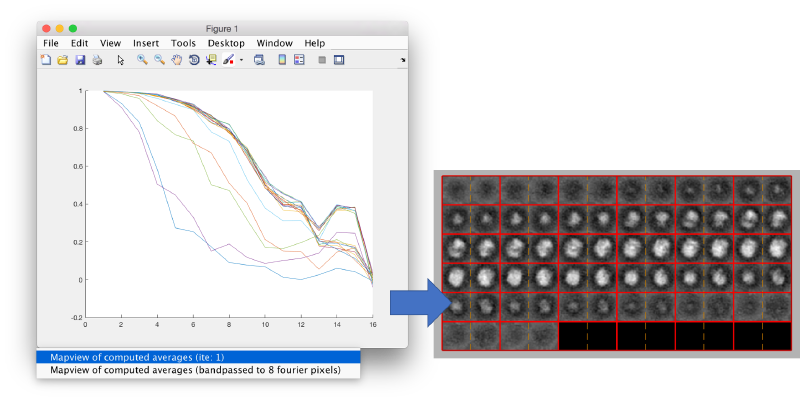
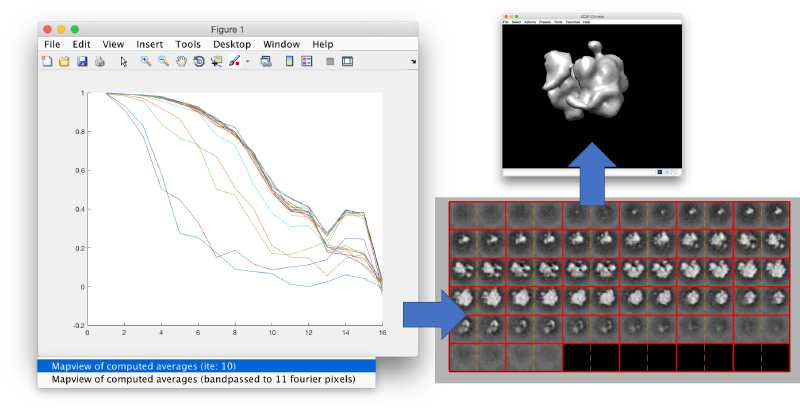
You can left click on each curve to identify the iteration number and to show the attained average.
Creating an ABP project from the command line
We show now how to quickly create an ABP project from the command line.
Create the support project
dcp.new('ptest02','d','ttest2/data','show',0);
Provide initial files
The rest of the seed files can initialized to default values through
dvhint_files ptest02 item *;
Change execution parameters
Numerical parameters can be introduces through dvput
dvput ptest02 -cr 360 -cs 45 -ir 360 -is 45 -rff 2 -rf 4 -dst matlab_parfor -cores 4 -mwa 4 -dim 16 -ite 10
The first six define the angular parameters of the project.
- dst defines the execution environment (matlab_parfor or standalone)
- cores number of cores during alignment.
- mwa number of cores during average.
Derive ABP project
Now the corresponding ABP project can be derived:
dveo ptest02 --abf
By default, the derived project will be called ptest02_eo. The tables are automatically split into two sets of particles. The templates can be created and introduced into the project with:
dvtemplate ptest02_eo --ro -N 10
We can compute the FSC of the constructed templates:
dpkproject.fsc.plotFSCInitialReferences(dvload('ptest02_eo'));
Configure ABP project
Now we can pass the specific parameters for adaptive bandpassing:
dvput ptest02_eo -abpi 3 -abpt 0.143 -abppb 0 -ite 10 ite_r2 0 -ite_r3 0 -lim 3 -limm 1
- abpi is the initial bandpass parameter (in Fourier pixels)
- abpt is the bandpass threshold
- abppb is the pushback parameter.
Now, we only need to configure the masks to default values.
dvhint_files ptest02_eo item masks
And we are ready to check:
dvcheck ptest02_eo
unfold:
dvunfold ptest02_eo
and run
ptest02_eo
or, in the standalone:
./ptest02_eo.exe
The results can be checked through:
dpkconv.plotConvergenceAll('ptest02_eo');
The resolution in Fourier pixels for each step is in the database of the project and thus accessible through ddb
ddb ptest02_eo:bandpass_resolution -r t
The content of the variable t can be displayed through:
disp(t);
Extending an ABP project from the command line
We can extend a project to continue refining previously computed results:
dynamo_vpr_branch ptest02_eo refine02 -n 0 -ite 7 -br 1
The new project refine02 has the same angular parameters as the original ptest02_eo. We change them to produce a local, finer grid.
dvput refine02 -cr 15 -cs 3 -ir 15 -is 3 -rff 2 -rf 2 -ite 10 -abpi 8 -mra 0 -dim 32
Note that we also
- use unbinned particles -dim 32
- start with a less conservative initial bandpass.
The project can now be unfolded
dvunfold refine02
and executed.
refine02
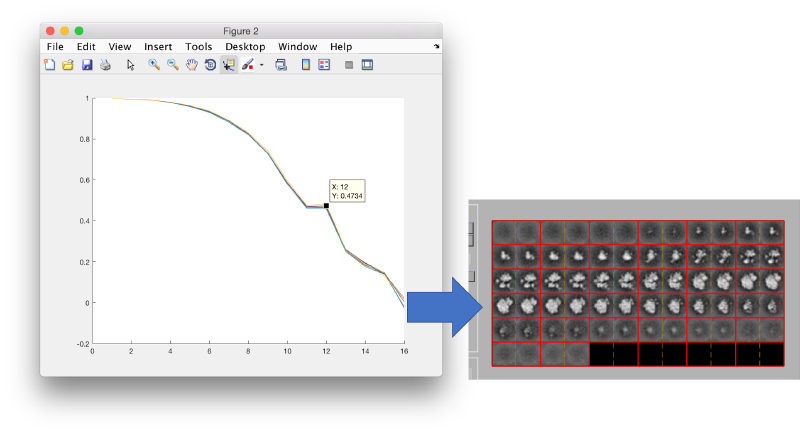
Upon completion, the iteration history can be seen through.
dpkconv.plotConvergenceAll('refine02');
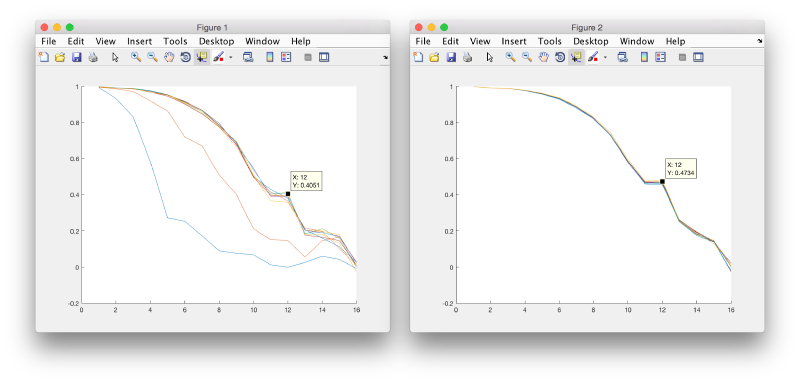
When comparing the results with the previous project, although there is no increase on the maximal bandpass attained, we can check that the FSC is larger for the same frequencies.