Walkthrough on command line based tilt series alignment
Dynamo includes a package for automated alignment and reconstruction of tilt series. This walkthrough guides users through a series of steps on how to use it in command line based mode. The GUI based alignment walkthrough shows how to operate this procedure through the GUI. Here, we go through each command one by one (for advanced applications, all commands can also be summarized in a Matlab script and run at once).
Contents
Create the workflow
The following commands create a new alignment workflow in a separate folder. Make sure Dynamo is running and type the following commands into the Matlab command line (or the Dynamo console, if you work on the standalone version):
name = 'hivCommandLine'; folder = 'workflows'; u = dtsa(name,'--nogui','-path',folder,'fp',1);
In the Matlab workspace you should now see the variables that you just created with these commands. If you work with the standalone version you can simply type the word whos to see a list of all variables in the workspace.
Entering the data
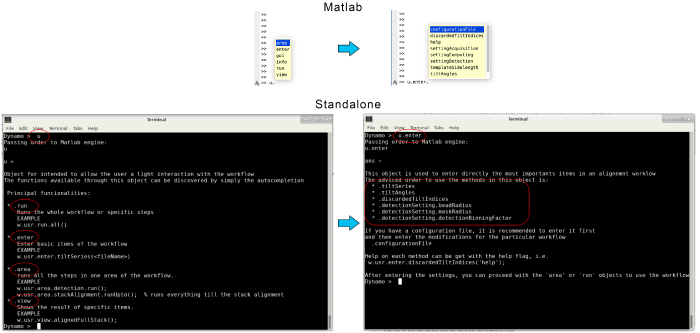
The variable u is an object that contains several fields to interact with the workflow. They can be found through autocompletion by using the tab key (or by pressing the enter key in the standalone version). In the following, these different fields will be used to pass our inputs to the workflow.
Basic data
Basic data can be entered using the enter field. Here we use a binned version of the first tilt series from the EMPIAR entry 10164, depicting a set of virus like particles (VLP). Visit the course materials to access the tilt series or download it directly from EMPIAR (you can also use your own data). The file of the tilt series can be first defined in a normale variable as
file = 'b001ts001.mrc';
and then passed to the workflow together with the angles:
u.enter.tiltSeries(file); u.enter.tiltAngles(-57:3:60);
If necessary (not in this tutorial), you can use the discardedTiltIndices field to reject tilts that are of too bad quality using the following command (e.g., tilt 1 and 40):
u.enter.discardedTiltIndices([1,40]);
Acquisition settings
Here you can enter parameters such as Cs, nominal defocus, etc. For this tutorial we just provide the pixel size in Angstroms:
u.enter.settingAcquisition.apix(2.7);
Computation settings
We enable the use of parallel cores
u.enter.settingComputing.parallelCPUUse(1);
This will be using mainly for gold bead detection (cc computation) and reconstruction.
To use all available physical cores use:
u.enter.settingComputing.cpus('*');
Detection settings
These here are the actual design decisions when running an alignment workflow. The gold bead radius has to be known beforehand ( see GUI based alignment walkthrough on how to measure it). Binning is used internally only for quick and robust bead detection.
u.enter.settingDetection.detectionBinningFactor(1); u.enter.settingDetection.beadRadius(16); u.enter.settingDetection.maskRadius(28); u.enter.templateSidelength(64);
Changing generic parameters
You can find all parameters of the individuals steps using autocompletion on the area field and then by adding step in the end. For example, the residual threshold of the tilt gap filler can be changed as follows:
u.area.indexing.step.tiltGapFiller.parameterSet.residualsThreshold(8);
Running the workflow
After setting the input parameters, the complete workflow can be run using the following command (we skip the CTF correction here). It should take about 5 minutes to run.
u.run.all('noctf',1);
Checking the results
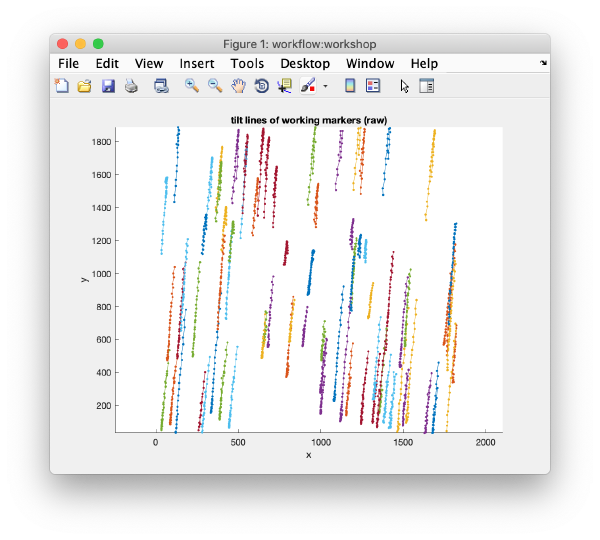
After completion, the view field allows you to check different output items for diagnose, as the tilt lines:
u.view.tiltLines.unaligned();
or the reconstructions, as for example
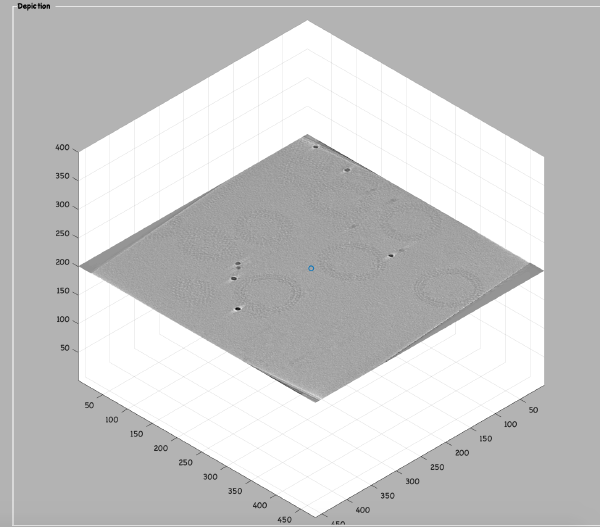
u.view.reconstruction.binnedWBP();
or
u.view.reconstruction.binnedSIRT();
and aligned stacks themselves.
u.view.stack.alignedFull();
You can also use the autocompletion (in the Matlab version) of the info field in order to read the logs on the fit.
>> u.info.fit; ------------------------------------------------------------ File with info: workflows/workshop2.AWF/info/fitting.doc - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - rms: 1.39 psi: 85.10 fit modus: normal total markers: 78 total observations: 2590 fit created at: 27-Aug-2019 18:01:40 ------------------------------------------------------------
or of the markers
>> u.info.markers; ------------------------------------------------------------ Info on working markers [used during detection, indexing refinement] ... item saved and in memory Item in memory: Number of shapes : 78 Number of frames : 40 Number of observations : 2590 (out of 3120) Number of empty frames : 2 Number of complete frames : 0 Number of complete shapes : 0 Item in disk: Markers in disk and currently in memory contain equal coordinates - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - Info on final markers [used from alignment step] ... item saved and in memory Item in memory: Number of shapes : 78 Number of frames : 40 Number of observations : 2590 (out of 3120) Number of empty frames : 2 Number of complete frames : 0 Number of complete shapes : 0 Item in disk: Markers in disk and currently in memory contain equal coordinates ------------------------------------------------------------
On the fly reconstruction of particles
In this section of the a, we show how to avoid reconstructing full tomograms of full resolution if only a set of subtomograms need to be reconstructed at high resolution. We will use the gold beads as proxies for particles of interest. We will thus locate a bunch of them in the binned tomogram, and then use their coordinates in that tomogram to directly reconstruct them in full resolution from the full resolution stack.
Grep coordinates in the binned tomogram
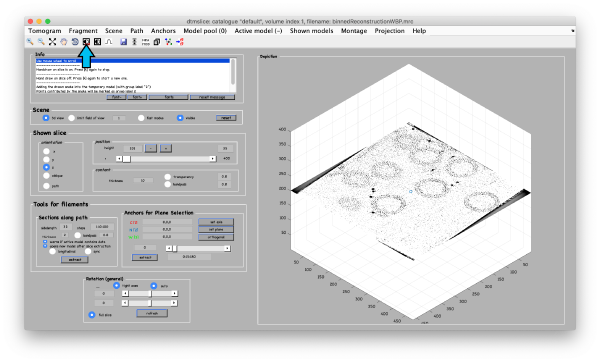
Open the binned reconstruction with dtmslice
dtmslice 'workflows/hivCommandLine.AWF/reconstruction/binnedReconstructionWBP.mrc';
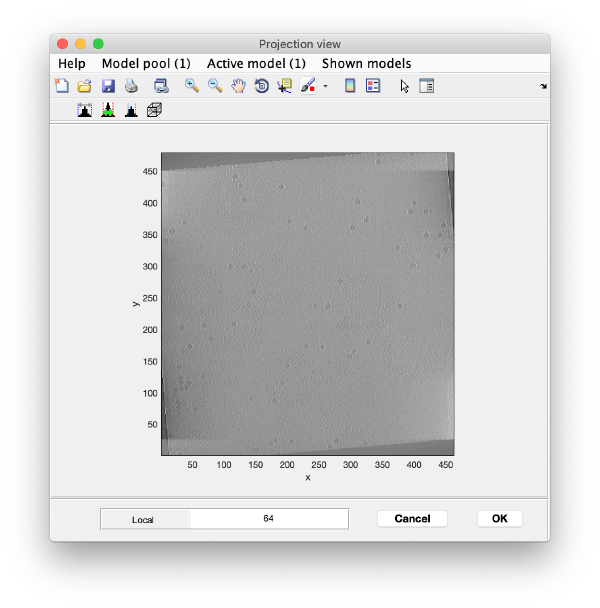
You will need to adjust the contrast:
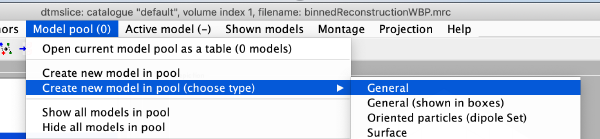
Now we create a new model of general type
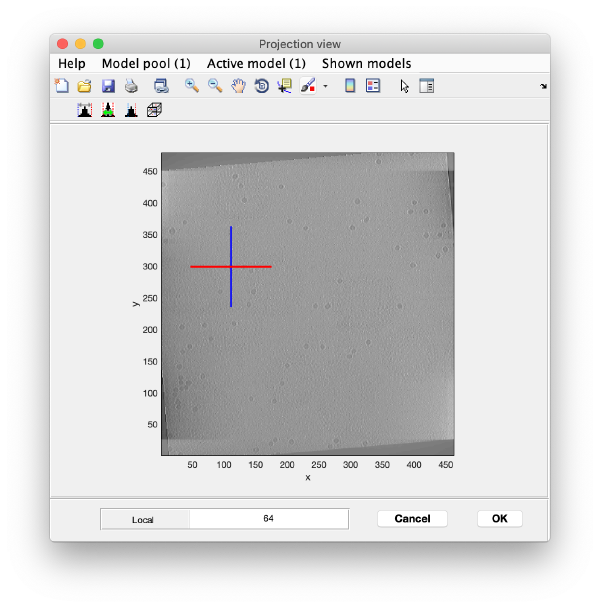
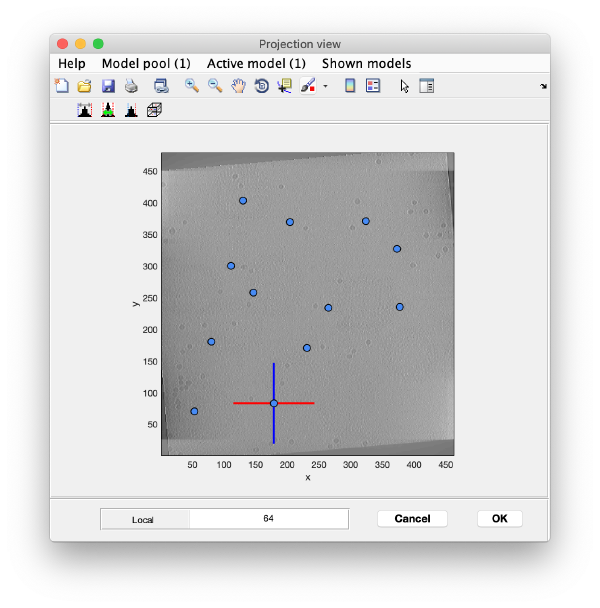
We use the projections in order to quickly find the 3d positions of a set of tomograms
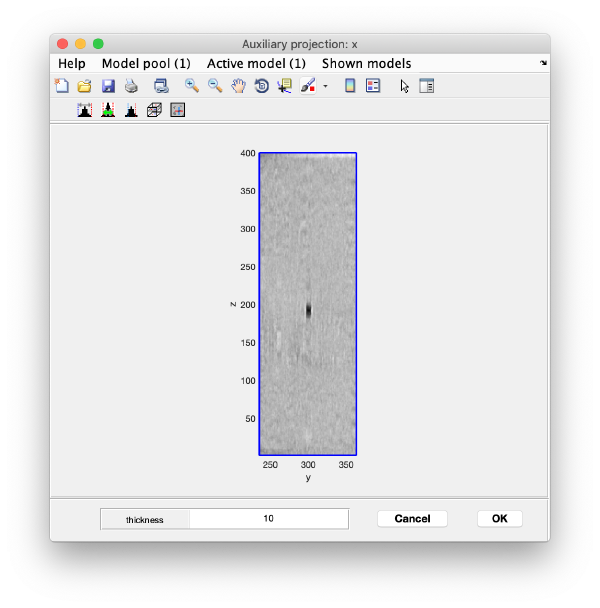
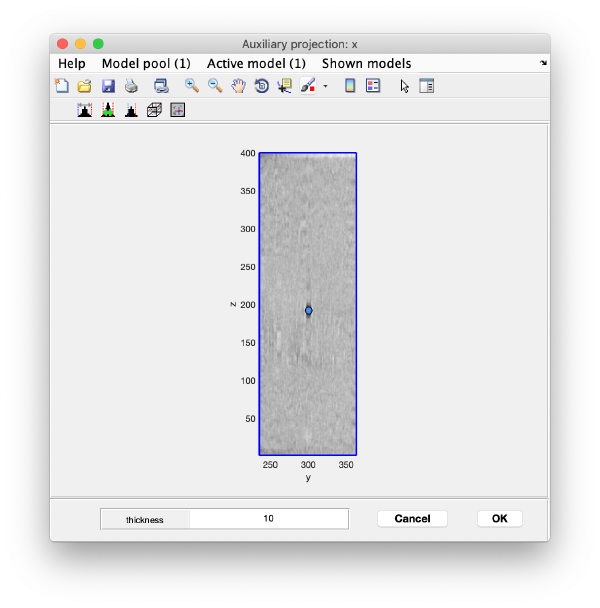
You can right click on the (x,y) position of a marker and get it depicted on its cutting xz and yz slices
Left clicking on it will provide a z coordinate to it
Keep clicking till you have a small set (no more than ten)
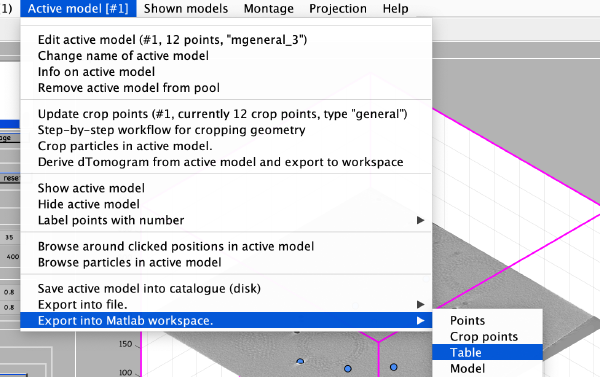
We export the coordinates of the model into the Matlab workspace already in table format, so that we will be able to use it in dtcrop on the binned tomogram.
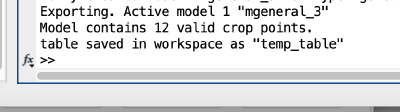
The Matlab shell should report the variable name used to store the table.
Now we can save the table as a file: write(temp_table,'binnedGoldbeads.tbl');
Cropping the gold bead particles
We can now crop the particles using the table file
tomo = 'workflows/hivCommandLine.AWF/reconstruction/binnedReconstructionWBP.mrc'; tbl = 'binnedGoldbeads.tbl'; targetBinnedParticles = 'binnedGoldBeads.Data'; sidelengthBinned = 32; o = dtcrop(tomo,tbl, targetBinnedParticles, sidelengthBinned);
And then we visualise a projection along z of each cropped particle
dslices(targetBinnedParticles,'proj','c');shg;
</nowiki>
Reconstruct particles on the fly
The command that reconstructs full sized particles from an aligned stack is dynama_table_trec. This is an independent function, not part of the workflow, implying that we will need to perform some format conversion.
tbl = dread('binnedGoldbeads.tbl');
We scale the table. "Bin level 2" means that each pixel in the binned tomogram is 2 times twice its size in the full resolution tomogram. This is the factor that we input into dynamo_table_rescale:
alignmentBinLevel = 2; tblFull = dynamo_table_rescale(tbl,'factor',2^alignmentBinLevel);
Now we fetch the full sized aligned stack stored inside the alignment folder:
source = 'workflows/hivCommandLine.AWF/align/alignedFullStack.mrc';
We create a data folder to contain the particles to be reconstructed on the fly:
targetDirectory = 'tempCrop.Data';
Angles actually used during the reconstruction are contained in the reconstructionTiltAngles' item of the output folder. Note that they might differ from the nominal angles, as some views might have been eliminated, both explicitely by the user, of by the program not being able to align all tilts.
angles = 'workflows/hivCommandLine.AWF/align/reconstructionTiltAngles.tlt';
The (binned) coordinates were measured in a binned tomogram, whose dimensionality is requested by dynamo_table_rec. We need to provide dynamo_table_rec with
visualizationTomogram = 'workflows/hivCommandLine.AWF/reconstruction/binnedReconstructionWBP.mrc'; sizeTomogram = dynamo_read_size(visualizationTomogram); sizeTomogramFull = sizeTomogram*2^alignmentBinLevel;
The tomogram center can be shifted in order to account for a possible shift in the centre in the binned reconstruction that was used to crop the particles.
shiftTomogramCenter = [0,0,0];
We proceed now to the reconstruction. Our sid length should be 4 times the size we used before:
sidelengthFullSize = 128;
We use the applyRampFilter option in order to explicitely estate the kind of filter that we want during the back projection.
dynamo_table_rec(tblFull,source,angles,targetDirectory, sidelengthFullSize,....
'applyRampFilter',1,....
'sizeTomogram',sizeTomogramFull,....
'shiftTomogramCenter',shiftTomogramCenter);
We can now check that the particles have been reconstructed in full size:
figure;dslices(targetDirectory,'proj','c');shg;