Difference between revisions of "Walkthrough for lattices on vesicles"
| Line 23: | Line 23: | ||
[[ HivDtmshow.png |thumb|center|500px| <tt>dtmshow</tt> on the sample tomogram. ]] | [[ HivDtmshow.png |thumb|center|500px| <tt>dtmshow</tt> on the sample tomogram. ]] | ||
| − | == | + | == Annotation of a tomogram == |
We are interested in cropping the particles between the two layers of each one of the viruses. | We are interested in cropping the particles between the two layers of each one of the viruses. | ||
| Line 44: | Line 44: | ||
Here, the user probably clicked on <tt>c</tt> for vesicle 2 in its real center, then clicked on <tt>n</tt> for the north point, and then moved to vesicle 3, clicking on <tt>c</tt> trying to mark the center. As a new dipole was not opened (i.e., the <tt>enter</tt> key was not pressed) when the vesicle 2 was finished, the center of vesicle 2 got misplaced. When this happens, simply press on <tt>h</tt> to hide the spheres and click the center of vesicle 2 back in its position. Then, press on <tt>enter</tt> to start the next vesicle. | Here, the user probably clicked on <tt>c</tt> for vesicle 2 in its real center, then clicked on <tt>n</tt> for the north point, and then moved to vesicle 3, clicking on <tt>c</tt> trying to mark the center. As a new dipole was not opened (i.e., the <tt>enter</tt> key was not pressed) when the vesicle 2 was finished, the center of vesicle 2 got misplaced. When this happens, simply press on <tt>h</tt> to hide the spheres and click the center of vesicle 2 back in its position. Then, press on <tt>enter</tt> to start the next vesicle. | ||
| + | |||
| + | When you are finished, don't forget to save the tomogram into the catalogue | ||
| + | |||
| + | == Cropping the particles == | ||
| + | |||
| + | We will describe a protocol to be carried from the command line, explaining each step. In short, we will visit each dipole in the model we just created, define a spherical vesicle centered on the dipole, use it to define a regular distribution of points on each vesicle, define positions pointing outwards and format everything as a [[table]] that can be used directly by ''Dynamo'' to operate an extraction and a subsequent alignment. | ||
| + | |||
| + | === Extracting information from the catalogue === | ||
| + | |||
| + | |||
| + | |||
| + | === Create a table for each vesicle === | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==== Merging the tables === | ||
| + | |||
| + | === Create a data folder === | ||
| + | |||
| + | === Create an average === | ||
| + | |||
| + | === Recroping the particles == | ||
| + | <tt>help dmodels.dipoleSet.getTableFromEnclosingSpheres</tt> | ||
Revision as of 17:03, 26 July 2017
This walkthrough shows the approach to process proteins that cover spheroidal vesicles following (approximately) a structured lattice. For demonstration, we use a highly binned tomogram with several viral capsides.
Contents
Data
The data has been made available by Florian Schur . This single tomogram is part of the data set used for the paper:
Getting the data
You can download the data through the command:
dpkhelp.wiki.downloadExample('hiv');
In case this command files, please try the direct link:
wget to be filled XXX
Viewing the data
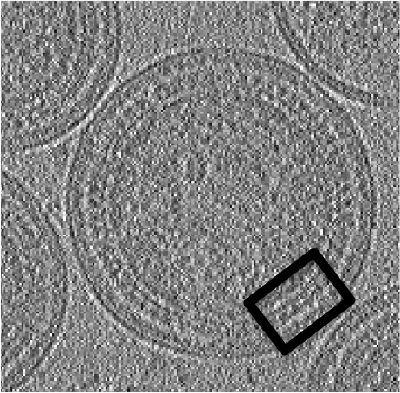
The tomogram is severely binned, so it will probably fit in memory without major problems. We can just use our typical tool to quick check a tomogram.
dtmshow v17.rec
thumb|center|500px| dtmshow on the sample tomogram.
Annotation of a tomogram
We are interested in cropping the particles between the two layers of each one of the viruses.
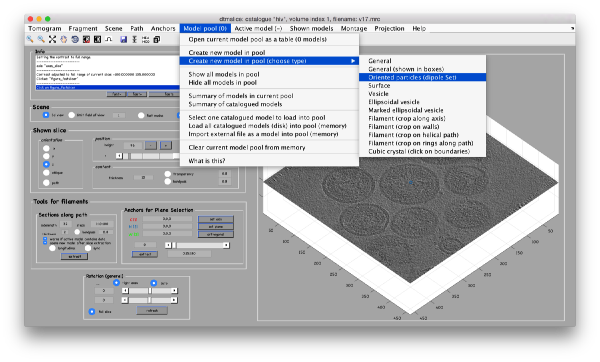
In our approach, we will first use a single model to manually input the approximate centers and radiuses of all the viruses. We will use the model type dipoleSet. This model allows to describe each annotated virus as a dipole: we will mark the (approximate) center of the virus as the center property of a dipole. The radius of the virus will be described by an annotation of the north property of the dipole.
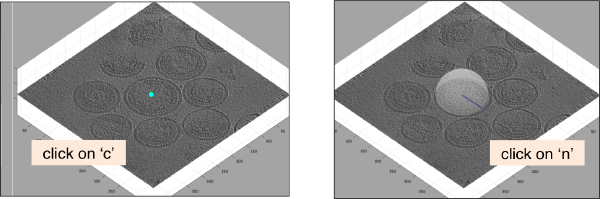
When the model is active, we use the key c to mark the center of the current dipole, and n to mark the north. It will create a semitransparent sphere enclosing the completed dipole.
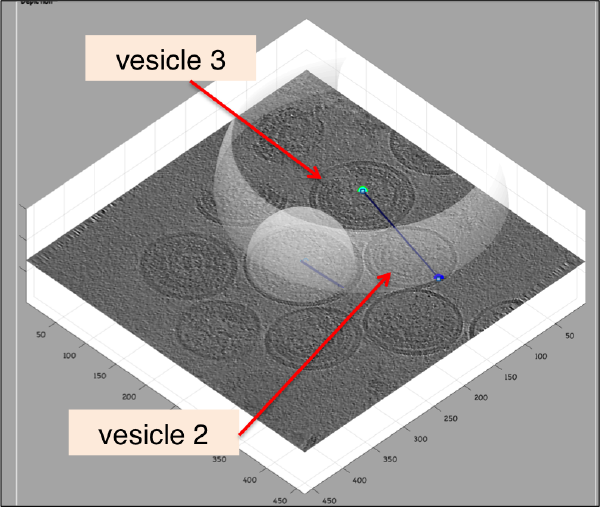
Further clicks on c or n will just move the center or the north of the same dipole. In order to click a further dipole, you need to click on enter and "open" the next dipole for clicking. If you forget to click on enter before annotating points on the next vesicle, you'll end up with situation like this:
Here, the user probably clicked on c for vesicle 2 in its real center, then clicked on n for the north point, and then moved to vesicle 3, clicking on c trying to mark the center. As a new dipole was not opened (i.e., the enter key was not pressed) when the vesicle 2 was finished, the center of vesicle 2 got misplaced. When this happens, simply press on h to hide the spheres and click the center of vesicle 2 back in its position. Then, press on enter to start the next vesicle.
When you are finished, don't forget to save the tomogram into the catalogue
Cropping the particles
We will describe a protocol to be carried from the command line, explaining each step. In short, we will visit each dipole in the model we just created, define a spherical vesicle centered on the dipole, use it to define a regular distribution of points on each vesicle, define positions pointing outwards and format everything as a table that can be used directly by Dynamo to operate an extraction and a subsequent alignment.
Extracting information from the catalogue
Create a table for each vesicle
= Merging the tables
Create a data folder
Create an average
= Recroping the particles
help dmodels.dipoleSet.getTableFromEnclosingSpheres