Difference between revisions of "Walkthrough for lattices on vesicles"
(Created page with "This walkthrough shows the approach to process proteins that cover spheroidal vesicles following (approximately) a structured lattice. For demonstration, we use a highly binne...") |
|||
| Line 21: | Line 21: | ||
We are interested in cropping the particles between the two layers of each one of the viruses. | We are interested in cropping the particles between the two layers of each one of the viruses. | ||
| + | |||
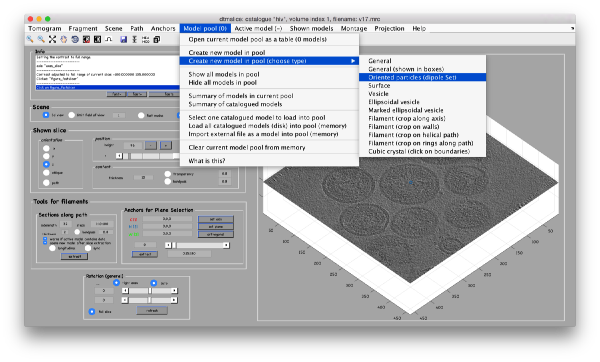
| + | In our approach, we will first use a single model to manually input the approximate centers and radiuses of all the viruses. We will use the model type <tt>dipoleSet</tt>. This model allows to describe each annotated virus as a ''dipole'': we will mark the (approximate) center of the virus as the <tt>center</tt> property of a dipole. The radius of the virus will be described by an annotation of the <tt>north</tt> property of the dipole. | ||
| + | |||
| + | [[File:HivSelectModel.png|thumb|center|600px| Selecting the ''dipoleSet'' model ]] | ||
| + | |||
| + | When the model is active, we use the key <tt>c</tt> to mark the <tt>center</tt> of the current dipole, and <tt>n</tt> to mark the north. | ||
| + | |||
| + | [[|thumb|center|600px| Selecting a ''dipole'': click on <tt>C</tt> for the center, then <tt>N</tt> for the north]] | ||
| + | |||
| + | Further clicks on <tt>c</tt> or <tt>n</tt> will just move the | ||
Revision as of 15:44, 26 July 2017
This walkthrough shows the approach to process proteins that cover spheroidal vesicles following (approximately) a structured lattice. For demonstration, we use a highly binned tomogram with several viral capsides.
Data
The data has been made available by Florian Schur . This single tomogram is part of the data set used for the paper:
Getting the data
You can download the data through:
XXXXXXX
Viewing the data
The tomogram is severely binned, so it will probably fit in memory without major problems. We can just use our typical tool to quick check a tomogram.
dtmshow v17.rec
Particle cropping
We are interested in cropping the particles between the two layers of each one of the viruses.
In our approach, we will first use a single model to manually input the approximate centers and radiuses of all the viruses. We will use the model type dipoleSet. This model allows to describe each annotated virus as a dipole: we will mark the (approximate) center of the virus as the center property of a dipole. The radius of the virus will be described by an annotation of the north property of the dipole.
When the model is active, we use the key c to mark the center of the current dipole, and n to mark the north.
[[|thumb|center|600px| Selecting a dipole: click on C for the center, then N for the north]]
Further clicks on c or n will just move the