Viewing tomograms
Dynamo offers several ways to visualize tomograms. The choice of one or other program depends on varios factors, like "How big is the tomogram?" "Are you going to annotate models on them?".
Contents
Tomogram browsers
Most browsers can be invoked directly from the command line or from a GUI linked to a catalogue.
The browsers you are most likely to use are:
- dtmshow quick, simple browsing outside of the catalogue system
- dpreview quick inspection of a few slides of catalogued volumes, for sending areas to other browser
- dtmview annotation of isolated particles and membranes inside the catalogue system
- dtmslice annotation of filaments and membranes inside the catalogue system
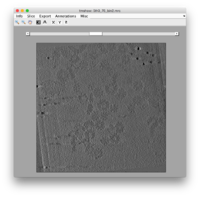
dtmshow
The simplest visualization system is the program dtmshow, callable from the command line and linked by many Dynamo GUIs. It's a lightweight slice-based browser. The user can choose to use it by preloading of the full tomogram or by accessing the disk for each depicted slice ("on-the-fly" modus). dtmshow does not allow to make complex annotations, or to register them inside the Catalogue via the model pool. Matlab documentation dtmshow
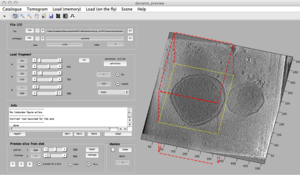
dpreview
dpreview goes a step further than dtmshow. It is connected with the Catalogue system, in fact 'open with dpreview' is one of the main options that you get when selecting a catalogued tomogram in the dcm GUI.
In dpreviewslices are always accessed "on the fly", so that browsing can be slow (especially if trying to depict slices orthogonal to the x- or y- directions, which demand multiple I/O operations to gather a slice whose pixels are scattered in the disk). The main function of the dpreview is to provide a "preview" of the disk contents of a tomogram, letting the user to specify and area of the tomogram and a binning level. This fragment will be read into memory and passed to dtmview or dtslice.
dtmview
- Main article: dtmview
dtmview is normally invoked from the dcm GUI, frequently using dpreview to select a fraction. It is oriented to show concurrently slices representing orthogonal cuts inside a volume with a high level of control of the number of depicted slices, and the part of the scene shown to the user.
The main utility of dtmview is for clicking sets of isolated particles.
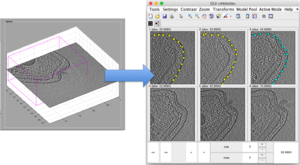
dtslicer
Is normally invoked by the dcm GUI or by dpreview . It depicts in 3d a slide or arbitrary orientation that can be moved freely inside the tomogram. This is a suitable browser for filament geometries. It is also suitable to locate the extent of membranes a send them over to the Sliding Montage browser for annotation.
Sliding montage
A sliding montage presents simultaneously a selection of slides of the tomogram. The user can choose to distance between consecutively depicted slices, and navigate inside the tomogram.
Sliding montages of areas inside a tomogram are specially useful for modelling membranes. Sliding montages are connected with the model pool and provide semiautomatic tools to track points manually picked in one level on the following levels.
Binned tomograms
Binning tomograms produces volumes much easier to fit in memory (1x binning ammounts to 8 times les voxels), and also clearer images. Thus tt is almost always useful to visualize binned versions of the original tomograms, even though the subtomograms for averaging will be cropped from the big, high resolution tomograms.
In all the volume browsers linked to the catalogue the idea is that you shouldn't care about coordinate transformations of annotations produced on different versions of one tomogram. Coordinates of models and particles clicked on binned versions are stored in disk with the correct dimensionality.
Binning on-the-fly
This is the easiest way: when opening a tomogram (or a tomogram fragment) in memory inside dtmslice or dtmview you would normally (although) not necessarily do it through the dpreview GUI. This GUI allows you to select the binning level of the fragment that will be depicted in the annotating browsers dtmslice or dtmview. In those annotation browsers, the coordinates that you marked will be referred to the unbinned image.
For instance, if you have a tomogram 4k x 4k x 1k voxels and choose to use a binning level of two in dpreview before sending it to, say, dtmslice, dtmslice will internally contain in memory a chunk of [1k x 1k x 250 voxels]. But whenever you mark a coordinate in dtmslice it will still refer to the 4k x 4k x 1k voxels.
Note however, that this system can be annoying: when you open a tomogram in dtmslice it can take minutes to complete the binning (Dynamo takes some time in accessing the disk for each slice). Thus, if you are going to visit a tomogram several times, it is a good idea to use the options for pre-binning explained below.
Prebinned tomograms
We advice the use of prebinned tomograms. Those are automatically generated by Dynamo. In the dcm GUI, click on Catalogue > created binned versions > <choose your factor>. This will go over all the tomogram files currently available in the catalogue, and will create a binned version of each one of them.
For a file called, say, <myPath>/someTomogramName.mrc, the catalogue will create another one called <myPath>/someTomogramName_catBinned2.mrc. Whenever binned versions of <myPath>/someTomogramName.mrc are requested by any browser, Dynamo will look if a file that follows this convention exists, and directly load it instead of computing the binning on the fly.
It will take some time so that it's best to let Dynamo alone for while till finishing this task. If you add new tomograms to the catalogue and rerun this option, the volumes that have already been binned will be skipped. Also, you can also bin the tomograms yourself. As long as they have the right naming convention described above, the Catalogue will find them.
Explicite volume pairs
This the most complicated (but most flexible) system. It allows the user to enter in the catalogue two different files to represent one single tomogram. In the column file of the dcm GUI, you would introduce the name of a depiction volume. All created annotations will be based on the dimensions of this volume. Now, if you want to actually crop particles from a different volume, you need to enter it in the column cropping volume .
Once the pair is created, you can put a value of one in the column crop elsewhere . This way, each time you want to crop particles from this tomogram pair, the cropping function will access the tables stored for the depiction volume, scale them according to apix crop volume and shift crop volume and then crop the subvolumes from the cropping volume. Note that with this option, you need to specify the apix of the depiction volume itself.
An example script is provided in dpktomo.examples.catalogue.cropHighResPickLowRes, where you can follow how to define the parameter shift crop volume. This example also shows you how to use this functionality from the command line.
In this case, all model coordinates will be stored with the coordinates of the unbinned depiction tomogram. When the catalogue is requested to crop subtomograms, it will automatically generate cropping tables with coordinates scaled to the size of the cropping tomogram.
This feature is also useful when you want to use tomograms of the same dimensionality but different appearance for picking. For instance, you want to use tomograms reconstructed with SART (typically providing a nice contrast) for depiction and particle clicking.
Thumbnails
Thumbnails are a minor functionality of the Catalogue. They are just agressively binned versions of the tomogram files (normally down to 40x40x40 pixels), so that a tomogram can be depicted quickly in a montage. Thumbnails are useful to visualize gallery views of your full sets of tomograms.
Slices
Sometimes you will want to depict singles slices in a Matlab graphic window, i.e., not inside a browser. This is useful when creating complex graphic material for presentations, publications, etc, as you can edit the created graphics with total flexibility.
Dynamo provides two Matlab classes for this task: the easy-to-use dSlice and the more flexible dpktomo.volume.slices.Slice. Use help to check their syntax from the command line.