Difference between revisions of "Vesicle models"
Jump to navigation
Jump to search
| Line 14: | Line 14: | ||
=== Separation === | === Separation === | ||
Given in pixels. | Given in pixels. | ||
| − | Defines the mean separation of the cropping positions to be defined in the vesicle surface. | + | Defines the mean separation of the cropping positions to be defined in the vesicle surface. When choosing this value, you should think on [[Seed oversampling|oversampling]] the actual distribution of particles that you expect. |
Revision as of 09:38, 22 April 2016
Vesicle models are appropriate when particles are evenly distributed on surfaces reasonably similar to spheres or ellipsoids. The model can estimate the center and radius provided points provided by the user in the visible part of the object, or the other way round, letting the user provided center and radius.
Contents
Important parameters
Radius
Given in pixels. Can be estimated or provided by the user.
Center
Given in pixels. Can be estimated or provided by the user.
Separation
Given in pixels. Defines the mean separation of the cropping positions to be defined in the vesicle surface. When choosing this value, you should think on oversampling the actual distribution of particles that you expect.
Command line
Here come some examples of creation and manipulation of vesicle models from the command line
Given user points
Given center and radius
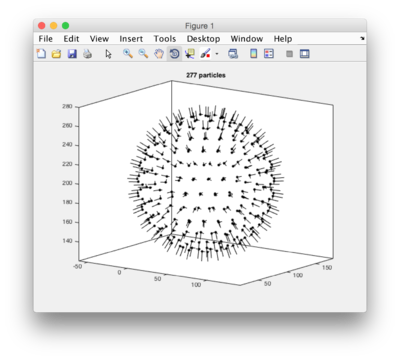
a = dmodels.vesicle(); % creates an empty vesicle model a.radius = 70; a.center = [40,100,200]; a.separation = 15; a.updateCrop(); % here it will some time computing the particles. t=a.grepTable(); % extracts the table dtplot(t,'profile','oriented_positions'); % scatter plot of images axis equal; % for a better visualization, same scale an x,y,z shg