Montage viewer for membrane picking
Revision as of 19:24, 27 February 2017 by Daniel Castaño (talk | contribs)
Here an example of use of the montage viewer for membrane models through the tomogram browser dtmslice is presented.

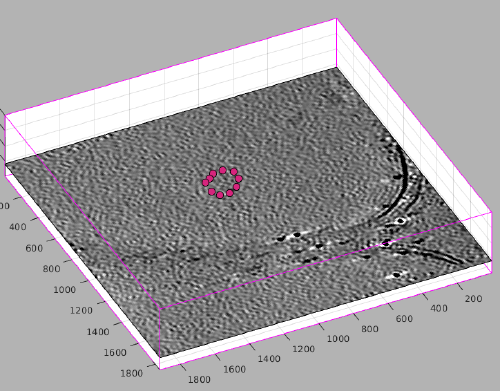
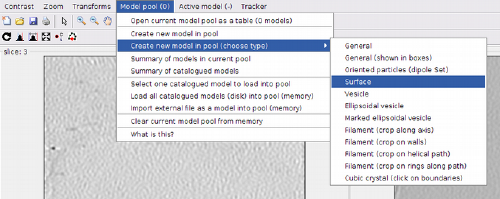
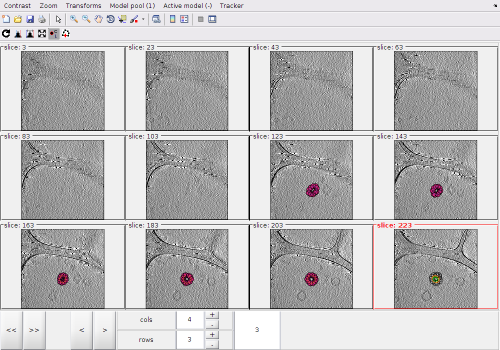
dtmslice view of a tomogram with several liposomes
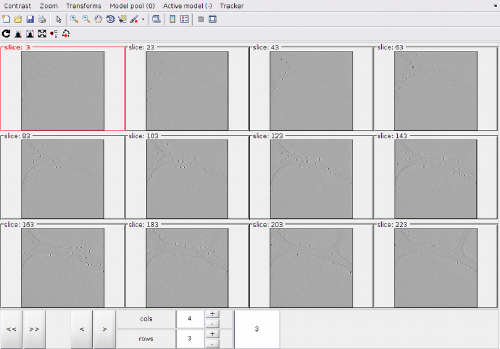
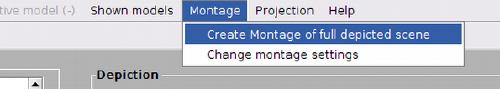
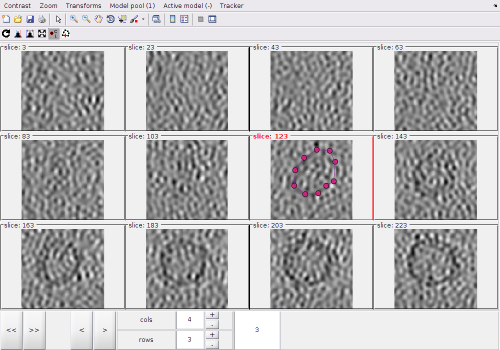
Montage view of the full dtmslice scene
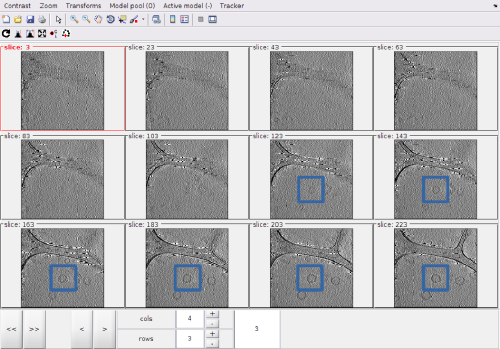
After activating the clicker, you can (main) click on a level of the montage.
The points created by clicking in the Montage Viewer will appear in dtmslice, and the other way round, as both viewers are connected to the same model pool
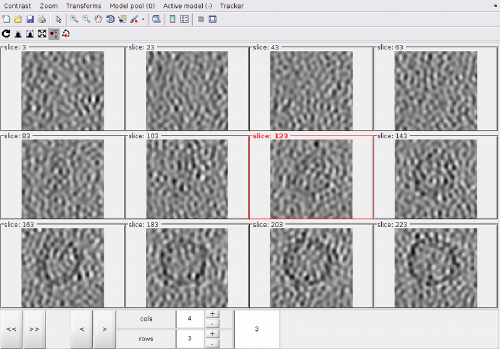
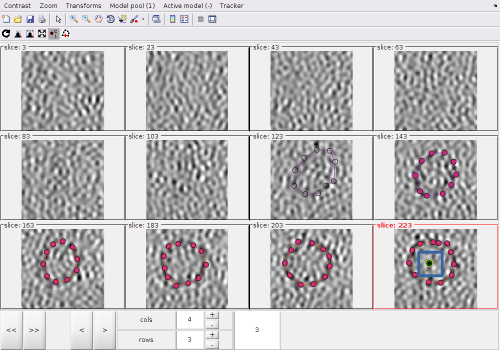
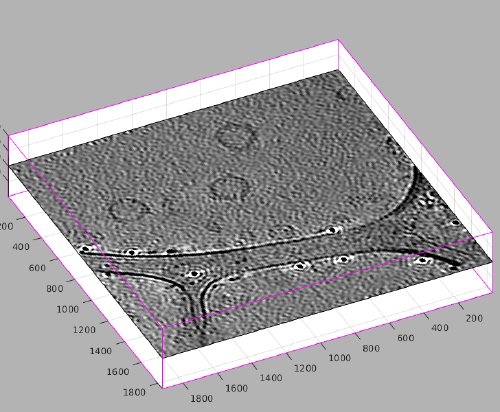
After visiting all the the zlevels where the membrane is defined, we can use the option for zooming out the view:
In this view, we can localte a new area of interest, create a new model and zoom again into the area of interest