Difference between revisions of "Montage viewer for membrane picking"
| Line 1: | Line 1: | ||
Here an example of use of the montage viewer for membrane models through the tomogram browser [[dtmslice]] is presented. | Here an example of use of the montage viewer for membrane models through the tomogram browser [[dtmslice]] is presented. | ||
| − | == Starting the Montage | + | == Starting the Montage Viewer == |
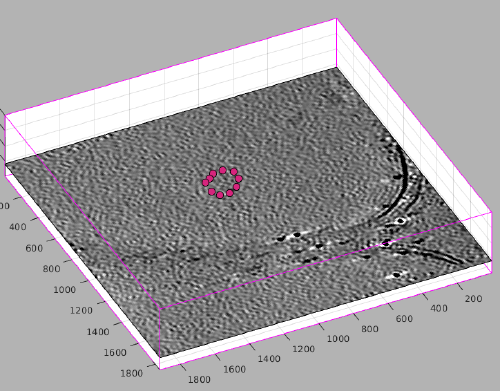
[[File:MontageMembranesTomogram.png|thumb|center|500px| [[dtmslice]] view of a tomogram with several liposomes ]] | [[File:MontageMembranesTomogram.png|thumb|center|500px| [[dtmslice]] view of a tomogram with several liposomes ]] | ||
| Line 7: | Line 7: | ||
[[File:MontageMembranesMenu.png|thumb|center|500px| Opening the Montage viewer for the full scene]] | [[File:MontageMembranesMenu.png|thumb|center|500px| Opening the Montage viewer for the full scene]] | ||
| + | |||
| + | === Options=== | ||
With the default options, the montage shows at each ''z'' level the full projection of the intensity between a level and the next one. | With the default options, the montage shows at each ''z'' level the full projection of the intensity between a level and the next one. | ||
[[File:MontageMembranesMontage.png|thumb|center|500px| Montage view of the full [[dtmslice]] scene ]] | [[File:MontageMembranesMontage.png|thumb|center|500px| Montage view of the full [[dtmslice]] scene ]] | ||
| + | |||
| + | === Model system=== | ||
The [[model pool]] currently accessed by <tt>dtmslice</tt> is also available through the Montage Viewer. Let us define a membrane model. | The [[model pool]] currently accessed by <tt>dtmslice</tt> is also available through the Montage Viewer. Let us define a membrane model. | ||
| Line 34: | Line 38: | ||
[[File:MontageMembranesZoomedFirstSelection.png|thumb|center|500px| Membrane points clicked on the selected ''z'' level ]] | [[File:MontageMembranesZoomedFirstSelection.png|thumb|center|500px| Membrane points clicked on the selected ''z'' level ]] | ||
| − | The points created by clicking in the Montage Viewer will appear in [[dtmslice]], and the other way round, as both viewers are connected to the same [[model pool]] | + | The points created by clicking in the Montage Viewer will appear in [[dtmslice]], and the other way round, as both viewers are connected to the same [[model pool]]. |
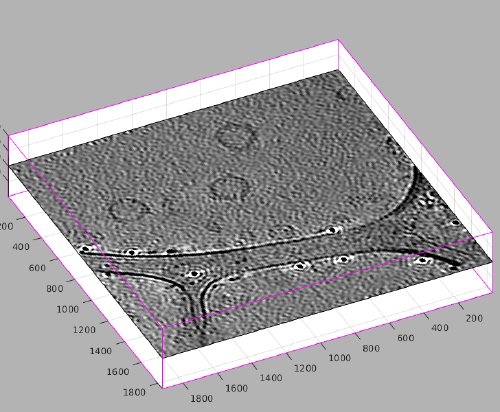
[[File:MontageMembranesLocateFirstScene.png|thumb|center|500px| The scene in <tt>dtmslice</tt> updates as we make annotations in the Montage Viewer]] | [[File:MontageMembranesLocateFirstScene.png|thumb|center|500px| The scene in <tt>dtmslice</tt> updates as we make annotations in the Montage Viewer]] | ||
Revision as of 19:59, 27 February 2017
Here an example of use of the montage viewer for membrane models through the tomogram browser dtmslice is presented.
Contents
Starting the Montage Viewer

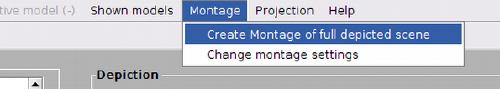
The Montage viewer can be opened from the menu options in dtmslice
Options
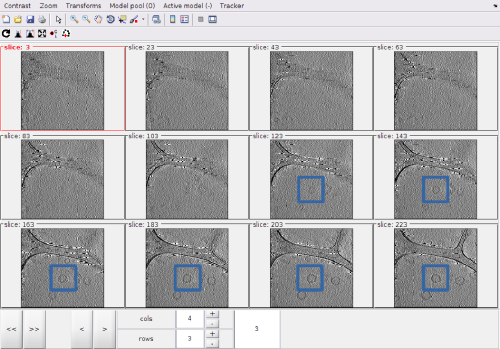
With the default options, the montage shows at each z level the full projection of the intensity between a level and the next one.
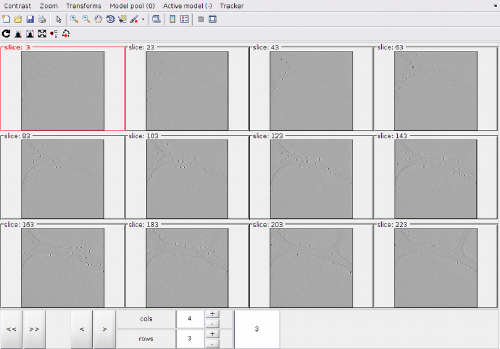
Model system
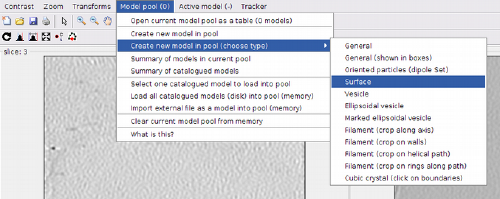
The model pool currently accessed by dtmslice is also available through the Montage Viewer. Let us define a membrane model.
Local zoom and drag
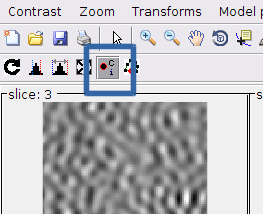
We make certain that the clicking option is active. This connects the tools for entering points into the model and to change the viewed scene in the montage (zoom and drag options).
With the full view, can inspect the scene and locate areas of interest.
After selecting the region of interest we can zoom on it, possibly also dragging the scene.
- Zoom in and out with the mouse wheel.
- Drag the scene by moving the mouse while the secondary button is pressed
- While the scene is being dragged, the cursor should appear as a hand.
Remember than the clicking tool must be activated in otder to have these controls operative:
Adding points
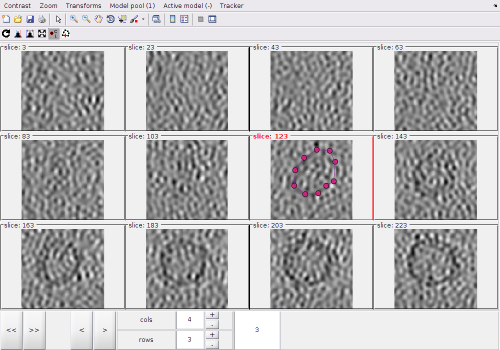
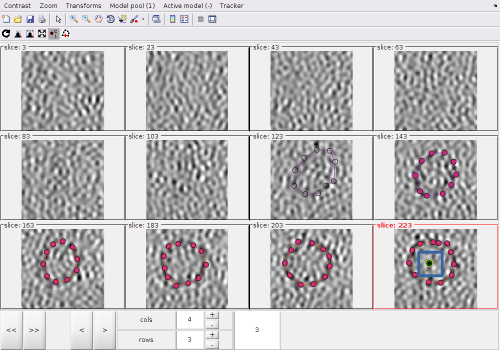
After activating the clicker, you can (main) click on a z-level of the montage. Each click creates a point.
The points created by clicking in the Montage Viewer will appear in dtmslice, and the other way round, as both viewers are connected to the same model pool.
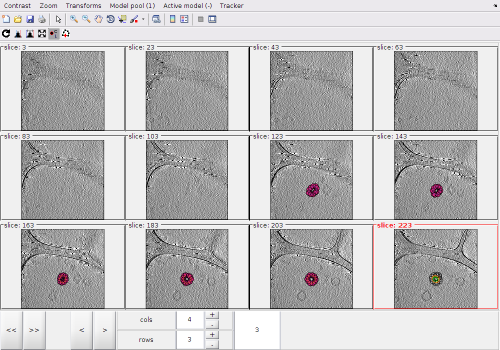
After visiting all the the zlevels where the membrane is defined, we can use the option for zooming out the view:
In this view, we can locate a new area of interest, create a new model and zoom again into the new area of interest