Getting a Structure from Multiple Tomograms of HIV Capsids (walkthrough)
In this walkthrough you will get familiar with managing multiple tomograms through the Dynamo catalogue and developing strategies for alignment projects to reach a reasonable structure from a set of tomograms. You will make use of the knowledge that you acquired during the workshop and apply it to a more realistic case. It is recommended to get familiar with at the advanced starters guide and the walkthrough for lattices on vesicles before starting this walkthrough.
Data
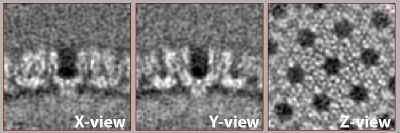
We prepared a set of 6 tomograms, each containing one Immature HIV-1 virus like particle(VLP) with a layer formed by a lattice of capsid proteins. The capsid proteins have a C6 symmetry and have a diameter of roughly 15nm and a molecular weight of about 150kDa. The pixelsize of the tomograms is 2.7 angstrom. You can find the tomograms in:
~/data/tutorial_VLPs
The tomograms are part of the data that was used in the publication An atomic model of HIV-1 capsid-SP1 reveals structures regulating assembly and maturation by Schur FK, Obr M, Hagen WJ, Wan W, Jakobi AJ, Kirkpatrick JM, Sachse C, Kräusslich HG and Briggs JA. (2016). The full dataset can be found on the Electron Microscopy Public Image Archive (EMPIAR) under EMPIAR-10164. The 6 tomograms were extracted from the tilt series TS_03 and TS_43.
Goal
From the data given you should be able to get a final average in which you start seeing hints of alpha helices, i.e., it should be possible to reach a resolution close to roughly 10 angstroms.
Walkthrough
Set up a catalogue
Create a new catalogue
dcm -create catVLP
and open it in the catalogue manager
dcm catVLP
You can add each volume manually to the catalogue under Catalogue -> Browse for new volume or you can make use of a volume file and add all tomograms at once under Input list of tomograms -> Browse for text file (.vll file). The volume file should be created beforehand by opening a blank text file named for example VLPtomograms.vll and copy the paths of all tomograms into it:
vlp_1.mrc vlp_2.mrc vlp_3.mrc vlp_4.mrc vlp_5.mrc vlp_6.mrc
In case you imported the tomograms using a volume file you have to click the button list volumes to refresh the tomogram list in the catalogue manager. Now you should see the 6 tomograms in the catalogue manager.
Define models
In each tomgoram you can now define
Open a catalogue
Add tomograms
Create vesicle models as described in
- Make sure you are familiar with the tutorial walkthrough for lattices on vesicles. Many of the described steps in the tutorial are needed in this exercise. You can also use similar parameters as used in the tutorial (adapted to the different pixelsize) for your first alignment projects.
- In the catalogue, you only need to prebin the tomograms with factor one (1x).
- Do not crop particles with a sidelength larger than 128px.
- Before using any C6 symmetries in the alignment projects, make sure to properly center the particles and template.
- To use the GPUs on sciCORE follow these steps.
- Without any postprocessing or elaborate refinement your final average may look similar to the one shown below: