EMBO workshop 2021
The tutorials listed here take place virtually on the 11th and 13th of September of 2021 (14:00-18:30) as part of the EMBO workshop for Image processing in cryoEM.
Contents
Connect to Guacamole and run Dynamo
Credentials
Please refer to your personal course material to access your two sets of credentials:
- A set of credentials to enter Guacamole (the platform for connecting via browser). These are common to all participants.
- A set of course credentials to enter the actual machines on which the course will be done: the EMBO21 UserID. These are unique to each participant.
Connect to Guacamole
To connect to Guacamole, first go to the following link in your browser:
https://login.cryst.bbk.ac.uk/guacamole/#/
At this stage you should maximize your browser window, otherwise you might have visualization problems when opening Dynamo. You should see the following window. Enter your Guacamole credentials.
Connect to your machine
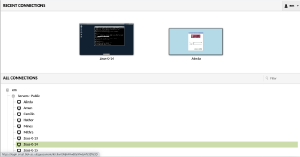
Now you need to select a machine. Please then click on +em and then +Servers - Public and select your dedicated server (in this example it is Zeus-0-14).
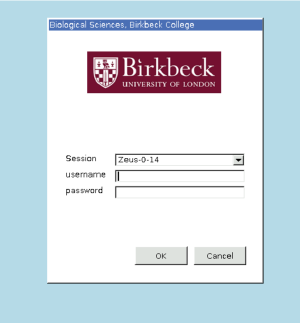
When you select the machine, you will prompted to use your EMBO21 UserID
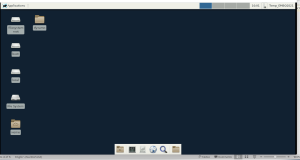
This should access your virtual Linux XFCe desktop, which should look like this in your browser
If you open a terminal (with the terminal icon) you can test by typing
pwd
that you are in the correct path, which should be:
/d/embo2021/u/emboXX
where XX stands for your actual workshop participant number.
Load and run Dynamo
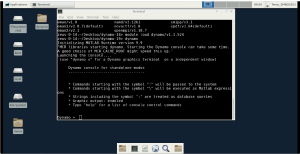
First you load the standalone version of Dynamo with the command:
module load dynamo/v1.1.524
and then it can be run by simply typing:
dynamo
Dynamo specific commands are then typed into this so called Dynamo console. For example, try to type the Dynamo command
dynamo_version
to see the current version that you are using. Note: Copy paste of commands from your local desktop into the dynamo standalone console on the remote Linux XFCe desktop might not always work. We suggest to additionally open the tutorials that you will be working on in a browser inside the remote Linux XFCe desktop itself, in case you need to copy paste long commands.
Type the following command to check how many CPU your machine has. You might use this number later for tasks that require parallel processing:
mbparse.multicore.checkPhysicalCores
Data location
The data used in the tutorials is already available for you. Move to the directory of the corresponding practical using the command:
cd prac-5
or
cd prac-6
respectively. Double-check if the data is present by typing
ls
You should see the following files in both directories:
During the Dynamo workshop, work only within those two directories.
Chimera
To use chimera, use the following command (replace 'myFile.mrc' with your file) in the normal terminal (outside Dynamo).
/usr/local/bin/chimera myFile.mrc
Schedule
The course is divided into short presentations and tutorials, as described in the time schedule below. For the presentations, we all meet in the main room. The tutorials are structured as follows:
- Demonstration: All students stay connected to the main room for the first minutes, where the goals of the tutorial are briefly explained.
- Individual work: Students go the assigned breakout rooms to work on the material independently. An instructor will be assigned to each breakout room to help with questions.
- Closure: During the last 5 minutes we come back to the general breakout room to wrap up the tutorial.
| Day 1 | |||
|---|---|---|---|
| Time | Activity | Topic | Comment |
| 14:00 - 14:30 | Presentation | Introduction | |
| 14:30 - 15:30 | Tutorial | Tilt series alignment with GUI | |
| 15:30 - 15:45 | Break | ||
| 15:45 - 16:45 | Tutorial | Tilt series alignment with command line | Skip last part "On the fly reconstruction of particles" |
| 16:45 - 17:00 | Break | ||
| 17:00 - 17:45 | Tutorial | Models for filaments | |
| 17:45 - 18:30 | Tutorial | Models for surfaces | |
| Optional: | Tutorial | Template matching | |
| Day 2 | |||
| Time | Activity | Topic | Comment |
| 14:00 - 14:15 | Presentation | Introduction | |
| 14:15 - 15:30 | Tutorial | Starters guide (workshop version) | |
| 15:30 - 15:45 | Break | ||
| 15:45 - 18:15 | Tutorial | Advanced starters guide | Take a 15 min break individually during this tutorial |
| 18:15 - 18:30 | Presentation | Closing remarks | |
| Optional: | Tutorials | Virus like particles or Spherical geometries | |