Difference between revisions of "Dynamo Workshop at the International School of Crystallography 2019"
Jump to navigation
Jump to search
| Line 1: | Line 1: | ||
This page describes the contents of the practical hands-on sessions for tomography through ''Dynamo'' in at the 54th International School of Crystallography in Erice. | This page describes the contents of the practical hands-on sessions for tomography through ''Dynamo'' in at the 54th International School of Crystallography in Erice. | ||
| + | A core presentation will be held on site | ||
== Computers == | == Computers == | ||
| − | === | + | === Download === |
| − | + | 'Information to be updated'' | |
=== Starting a ''Dynamo'' session === | === Starting a ''Dynamo'' session === | ||
| − | + | 'Information to be updated'' | |
| − | |||
== Program == | == Program == | ||
Revision as of 11:15, 13 February 2019
This page describes the contents of the practical hands-on sessions for tomography through Dynamo in at the 54th International School of Crystallography in Erice. A core presentation will be held on site
Contents
Computers
Download
'Information to be updated
Starting a Dynamo session
'Information to be updated
Program
Before the hands-on session, we will go to this presentation for a general introduction of the Dynamosoftware.
General Introduction

Clicking particles in the Starters guide
Guided presentation:
- tutorial on basic elements: help, data and metadata formats.
- tutorial on the basic concept in Dynamo alignment: the project.
Working on your own:
- Basic walkthrough: creating a catalogue, picking particles, launching a project.
- Advanced starters guide on a real tomogram. (~2 hours)
- Further work:
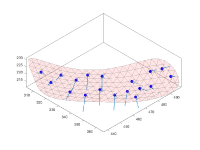
Geometric modeling
Short guided presentation:
- tutorial on membrane modeling with dmslice
- Filament models with dtmslice
- Reusing model workflows ( walkthrough)
- Further work: catalogue
Working on your own:
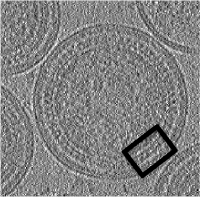
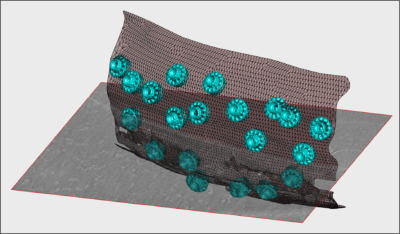
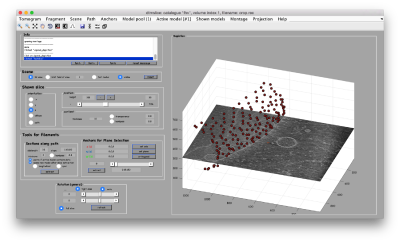
- In the afternoon, after the research talks, we will focus on the extraction of particles from densely packed spherical geometry on HIV viral capsides (~1 hour)
Template matching
Working on your own:
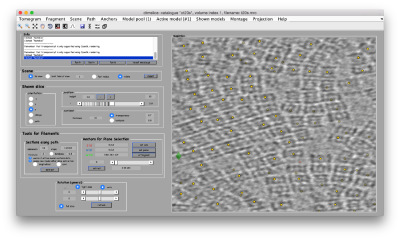
- We will follow this walkthrough for automated identification of proteosomes on a real tomogram through template matching. (~1 hour)
Adaptive bandpass filtering
Working on your own:
- We will follow this walkthrough to create a small synthetic data set that illustrates the principles of adaptive bandpass filtering, a way of conducting a golden standard alignment procedure . (~40 mins)
Classification
Short guided presentation:
- PCA Basic concepts.
advanced subboxing tutorial: combining with PCA and MRA tutorial - Multirefererence Analysis
Basic concepts.
walkthrough (~10 min) - Further work
- Commandline operations with PCA
Creation of 3D scenes
Working on your own:
- Walkthrough on the FHV data set (~1hour)
Further support material.
- Walkthrough on depiction and manipulation of triangulations (synthetic data).
Additional tools
Wednesday afternoon session.
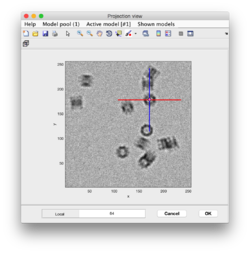
- Subboxing
- motivation slide: extraction of vertices from icosahedral viruses
- Basic subboxing (tutoriall)
- advanced subboxing tutorial: combining with MRA tutorial
- suggested data set: prd1 Capsides (in the folder of isolated particles)
- Manual alignment.
- Exercises
Data sets
The data sets will be updated here.
Instructors
- Daniel Castaño-Díez, University of Basel.