Delft Workshop 2017
A guide to use Dynamo during the TU Delft workshop for Cryo-EM. This covers the afternoon session of the 6th of September of 2017.
Preinstalling the software
We suggest students to download, install and test the software before departing for the workshop. Data should also be downloaded beforehand.
Downloading Dynamo
Download the package in these google links for linux or Mac respectively. After download (~4Gb), proceed to install and test
To install it, first create a folder in your filesystem in an arbitrary location of your choice:
mkdir <path to installation folder>
and untar the downloaded tarball into it:
tar -xf <tar file> -C <path to installation folder>
Testing your standalone installation
Activate your installation by typing in a Linux shell the activation script:
source <path to installation>/dynamo_activate_linux_shippedMCR.sh
or correspondingly for Mac:
source <path to installation>/dynamo_activate_mac_shippedMCR.sh
and then, to test the activated installation, just type in the terminal:
dynamo
You should get the Dynamo prompt.
Free license for Matlab
If the standalone installation works, you don't need to install further elements. However, we advice you to preinstall Matlab on your laptop. This makes the work with Dynamo much easier
You can get a trial version of Matlab and install it on your laptop. The license will be valid for 30 days. After installing Matlab, you can download the Dynamo version for your platform.
Testing your Matlab installation
Inside the Matlab shell type:
run <path to installation>/dynamo_activate.m
Virtual machines
If your system does not recognize the libraries provided in your package, you always have the option of installing a Virtual Machine in your system that emulates the Ubuntu distribution of linux. In this environment, Dynamo will run without errors, although the performance of the graphics system will be severely decreased. We recommend to use this device as a last resource.
Download an Ubuntu distribution
The version 16.04.3 of Ubuntu can be download as an iso file here
Download the virtual machine
A virtual machine can be downloaded here. A detailed walkthrough on the diverse steps can be found here
Data
Data should be downloaded beforehand:
wget https://wiki.dynamo.biozentrum.unibas.ch/w/doc/data/hiv/v17.rec wget https://wiki.dynamo.biozentrum.unibas.ch/w/doc/data/t20s/t20s.mrc wget https://wiki.dynamo.biozentrum.unibas.ch/w/doc/data/fhv/crop.rec
Accessing Delft servers
There is the possibility of connecting to a GPU machine of TU DELFT. The machine hpc04.tudelft.net has 8 GPUs installed.
Accessing the remote machine
For linux and Mac users, open a terminal on your laptop and type
ssh -Y <user>@hpc04.tudelft.net
User accounts will be distributed during the workshop.
Windows users do the same through their preinstalled virtual box.
Opening a Linux terminal in the remote machine
We need to create a command terminal. Right click on the screen and select Open Terminal in the menu . A Linux command window will popup. You can check the GPUS in the machine through writing:
nvidia-smi
Starting Dynamo
In the Linux terminal on hpc04.tudelft.net , we first need to make Matlab accessible invoking the corresponding module:
module load matlab
and then open it through:
matlab &
it will take some secons. Then, in the Matlab window write:
run <PATH TO DYNAMO> /dynamo/dynamo_activate.m
This will activate Dynamo in the current Matlab session.
Program
Introduction to tomography and subtomogram averaging
30 minutes of presentation to introduce tomography, subtomogram averaging and Dynamo.
Tutorial 1: Getting started
Morning session.
on screen presentation/hands-on practical. The presentation will loosely the introductory materials in:
- tutorial on basic elements: help, data and metadata formats.
- tutorial on the basic concept in Dynamo alignment: the project.
After the introduction of these general concepts, will follow together the basic hands-on tutorial of Dynamo.
Advanced getting started
In the afternoon session, participants will proceed to work on their own with the advanced introduction on a reduced data set from a real tomogram.
Further walkthroughs
Further walkthroughs are available for participants that complete this, they can be chosen according to the particular interests:
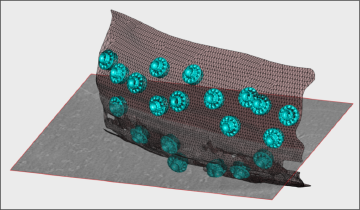
- Treatment of densely packed spherical geometry (~1 hour)
- walkthrough for automated identification of proteosomes on a real tomogram through template matching. (~1 hour)
- Walkthrough on the FHV data set (~1hour)
Instructors
- Daniel Castaño-Díez, BioEM Lab, University of Basel
- Paula Pérez-Navarro, C-CINA, University of Basel
- Cynthia Taveneau, Institut Curie, Paris