Construction of masks
Dynamo uses several types of masks in different pipelines.
Masks are used in Dynamo for several purposes: indicating the active part of the template for alignment, restricting the area of the aligned particles that will count for classification, avoiding FSC artifacts, placing aligned averages back in a tomogram....
In some tasks, the masks are expected to be simple geometrical shapes. In other occasions, it might be necessary to have a finer control on the process of constructing a mask. Dynamo includes several tools:
Contents
Geometrical Shapes
Use dmask to open a GUI with several geometrical options: spheres, ellipsoids, tubes, cylinders. dmask can be used from the command line.
There are other commands for specific geometries: dsphere, dshell, dcylinder....
Freehand draw
You can edit a template and draw on its 2D slices handdraw lines each enclosing an area of interest. There a different ways to convert such an area into a 3d volume
Revolution solids
Here, you would enclose an area on a 2d slice that includes the Z axis (i.e., a section from an X-view or a Z-view), and then let Dynamo rotate the area around the central Z axis to create a revolution solid.
This can be done with dynamo_mapview. Open your template, go to an X view, select one slice and press on [shift+S]. A second window will pop up, zooming into the selected slice. When you finish, the message area will tell you the name of the file containing the the created revolution solid.
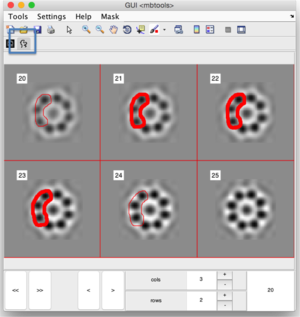
Slide by slide
You can edit a volume as a montage of orthoslices, and handdraw on each of them, or onto a selection of them an let Dynamo interpolate on the other sections:
dpkvol.montage('template.em');
Then choose the Mask tab to created a control for free handdraw of shapes on each slice. Secondary click on a drawn level will offer a menu of options, like interpolating between user-marked levels, or converting the current work into a cube.
By thresholding and smoothing
A mask can be defined to XXXXX
Combination of masks
It is easy to create combinations of masks using logical operators: or just "adds" masks.
Alternatively, you can use the compact syntax in dynamo_mask_combine on a cell array of masks defined independently. The flag delete can be used to define parts of the final mask that need to be set to zero (black).
Depicting mask overlays
When you design a mask, it is frequently useful to check how it relates to a given template. This can be done with the command dslices, using the flags overlay, overlay_as
For instance, if you have
dwrite(dynamo_sphere(16,64),'mask.em'); dslices template.em y -ov mask.em -ovas mask -ovc b