Considerations for tomogram positioning in IMOD
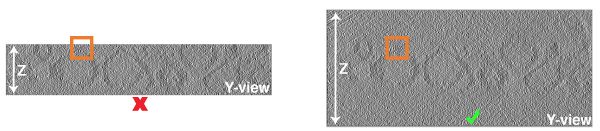
During tomogram reconstruction in IMOD at some point the dimensions of the final tomogram are calculated in the "tomogram positioning" step. In this step, we recommend to keep the x-axis tilt at zero and to leave enough room above and below the sample in the tomogram. Keeping the x-axis tilt at zero facilitates the interpretation of the tomograms, since the z-axis corresponds to the direction of the electrons. It also saves us from adapting the missing wedge mask at later steps. Leaving enough space at the top and bottom of the sample in the tomogram is necessary to crop subvolumes from sample that is close to the ice surfaces:
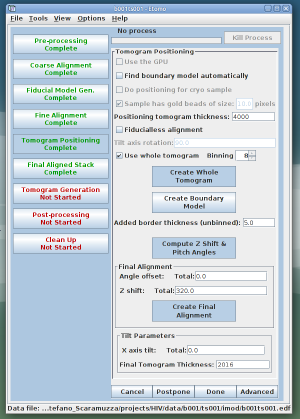
To keep the x-axis tilt at zero and to keep the tomogram thick enough we follow the following steps. First we choose a fairly large thickness for the sample tomogram (the sample tomogram will just be used to set the boundaries, it has not the final tomogram size). In the example below, an initial thickness of 4000 pixels was chosen:
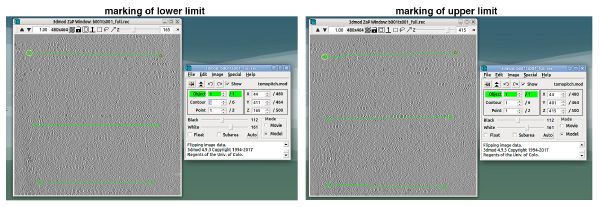
We also select a binning level of 4 or 8 to save computing time. After we create the sample tomogram we open it to set the boundaries. Here we do not follow the IMOD tutorial but just stay in the normal 3dmod view and set the lower and upper boundaries as shown below. This will prevent the introduction of any x-axis tilt. It is important that in this step we leave enough space above and below the sample. We should be very generous with that (as explained in the first figure). After saving the model and clicking on “compute Z shift & pitch angles” there should be displayed a new z-shift and final tomogram thickness but the x-axis tilt should be 0.0.