Difference between revisions of "Advanced starters guide"
| Line 42: | Line 42: | ||
Hereby, the flag <tt>-otf</tt> means "on the fly", telling <tt>dtmshow</tt> to ''not'' preload the full tomogram, but to access in disk the individual slices that are needed when inspecting a particular area. | Hereby, the flag <tt>-otf</tt> means "on the fly", telling <tt>dtmshow</tt> to ''not'' preload the full tomogram, but to access in disk the individual slices that are needed when inspecting a particular area. | ||
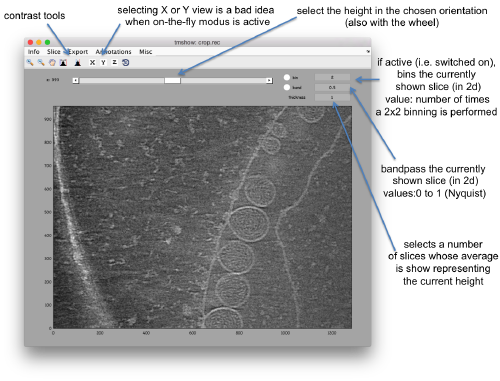
| + | [[File:FhvTomoshowOnTheFly.png|thumb|center|500px|Basic controls of <tt>dtmshow</tt>]] | ||
| + | |||
| + | Go up and down. We want to select the locations were the vesicles intersect the mythocondrion membrane and average them together. For this, we need to ''catalogue'' the tomogram, so that our annotations are stored with a clear relationship with the tomogram. | ||
| + | |||
| + | === Cataloguing the tomogram === | ||
| + | |||
| + | We can create catalogues just to contain a single tomogram. They are useful to keep track of all annotations, and of the typical transforms (binning, cropping of fractions) that we usually perform on a larged size tomogram of interest. In this case, we can create the catalogue directly from the command line: | ||
| − | + | <tt> >> dcm -c create fhv </tt> | |
| + | |||
| + | where <tt>fhv</tt> is just an arbitrary name. | ||
| + | >> dcm -c fhv -at crop.rec | ||
Revision as of 20:20, 15 July 2017
This walkthrough uses a small size example based on a real tomogram to covers several tasks.
Contents
The example data set
The data is a fraction of a tomogram. The full tomogram was used in "Cryo-electron tomography reveals novel features of a viral RNA replication compartment." (Ertel et al.), and represents several FHV viruses docked in the membrane of a mythocondrion.
Downloading
In principle, you can download all the files related to this example with the command:
dpkhelp.wiki.downloadExample('fhv');
If it fails under Matlab or the Dynamo command line, you can try to directly use the linux order
wget https://wiki.dynamo.biozentrum.unibas.ch/w/doc/data/fhv/crop.rec
or
curl -O https://wiki.dynamo.biozentrum.unibas.ch/w/doc/data/fhv/crop.rec
unter MacOS.
This should have created the file called crop.rec in your current directory.
Size check of a file
You're probably curious to see what's inside, so that let's write first:
dfile crop.rec
to let Dynamo check the dimensions of the file. The header of a .rec file is readed as a regular mrc, yielding:
filetype: volume size: 1285 x 956 x 786
So, it's a tomogram.
Lightweight visualization
We can inspect quickly its contents with
dtmshow -otf crop.rec
Hereby, the flag -otf means "on the fly", telling dtmshow to not preload the full tomogram, but to access in disk the individual slices that are needed when inspecting a particular area.
Go up and down. We want to select the locations were the vesicles intersect the mythocondrion membrane and average them together. For this, we need to catalogue the tomogram, so that our annotations are stored with a clear relationship with the tomogram.
Cataloguing the tomogram
We can create catalogues just to contain a single tomogram. They are useful to keep track of all annotations, and of the typical transforms (binning, cropping of fractions) that we usually perform on a larged size tomogram of interest. In this case, we can create the catalogue directly from the command line:
>> dcm -c create fhv
where fhv is just an arbitrary name. >> dcm -c fhv -at crop.rec