Difference between revisions of "Walkthrough for template matching"
| Line 60: | Line 60: | ||
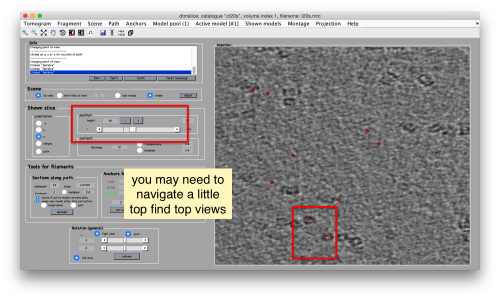
[[ File:TemplateMatchingTomoslicePickTop.png |thumb|center|500px| Use the wheel to go up and down in the tomogram to locate top views ]] | [[ File:TemplateMatchingTomoslicePickTop.png |thumb|center|500px| Use the wheel to go up and down in the tomogram to locate top views ]] | ||
| + | |||
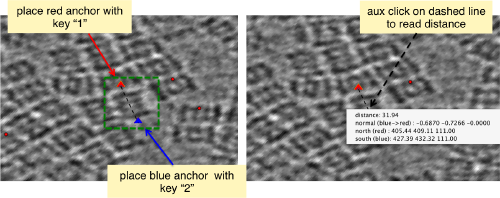
| + | [[ File:TemplateMatchingMeasure.png |thumb|center|500px ]] | ||
| + | |||
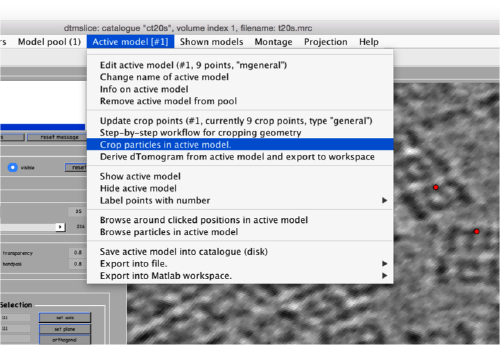
| + | [[ File:TemplateMatchingMenuForDtcrop.png |thumb|center|500px ]] | ||
| + | |||
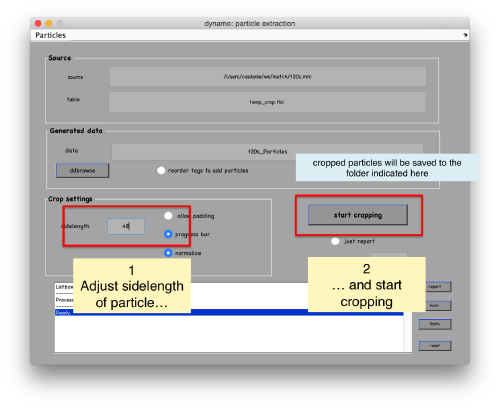
| + | [[ File:TemplateMatchingDtcrop.png |thumb|center|500px ]] | ||
| + | |||
| + | |||
====Manual alignment of some particles ==== | ====Manual alignment of some particles ==== | ||
| + | |||
| + | |||
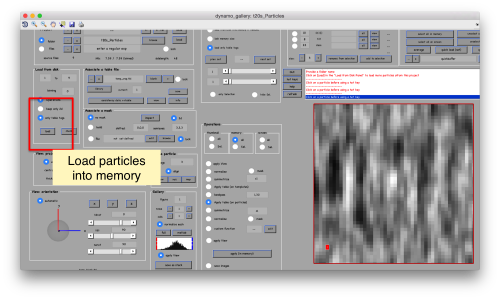
| + | [[ File:TemplateMatchingGalleryLoad.png |thumb|center|500px ]] | ||
| + | |||
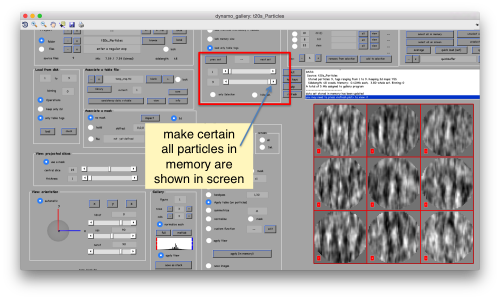
| + | [[ File:TemplateMatchingGalleryShowAll.png |thumb|center|500px ]] | ||
| + | |||
| + | [[ |thumb|center|500px ]] | ||
| + | |||
| + | [[ |thumb|center|500px ]] | ||
| + | |||
| + | |||
| + | |||
====Cropping the borders of a template==== | ====Cropping the borders of a template==== | ||
Revision as of 11:28, 19 August 2017
Dynamo includes a set of tools for location of particles inside tomograms. The most simple one is template matching.
Contents
Template matching
In this technique, a template representing a molecule of interest is systematically cross-correlated against a tomogram, producing a cross-correlation map of the tomogram. Each pixel in this map represents a score assigned the corresponding pixel in the tomogram map. This score measures the similarity of the neighbourhood of the tomogram pixel to the used template. This similarity is measured exclusively inside a mask.
Data set
Tomogram description
The tomogram contains a buffer with T20S proteasome on a holey carbon grid collected on a Krios + K2. Original pixelsize was 1.76 angstroms. The tomogram provided here has bin binned twice (yielding thus 7.04 ang), defocus is 4.4 microns, no CTF correction, no energy filter.
Acknowledgements
The tomogram has been kindly provided by Alex Noble, from the New York Structural Biology Center. Data collection was performed using Leginon and Appion-Protomo at the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy located at the New York Structural Biology Center, supported by grants from the Simons Foundation (349247), NYSTAR, and the NIH National Institute of General Medical Sciences (GM103310) with additional support from Agouron Institute [Grant Number: F00316].
Getting the tomogram
XXXXX
Visualizing the tomogram
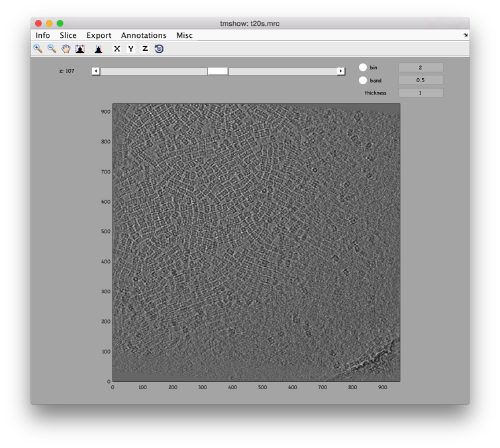
We can get a first glance on how the tomogram looks like:
dtmshow -otf t20s.mrc
to use the on-the-fly access to the slice shown at each given moment, or
v = dread('t20s.mrc');dtmshow(v);
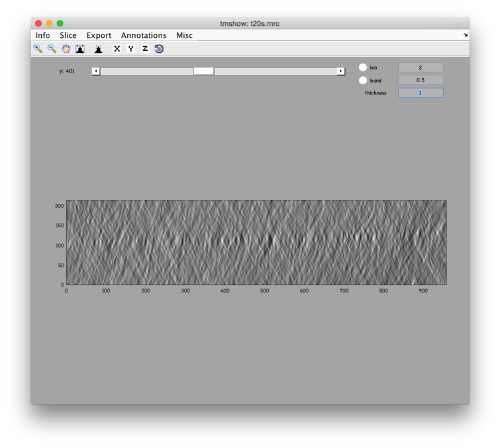
to preload the full tomogram into a memory variable (arbitrarily called v). In either option, you will see, the proteasomes are densely packed in an layer. The layer is slightly oblique, what can be seen browsing through z or y.
Navigating on-the-fly you'll see that transitions in y are slower than transitions in z, because all the pixels of the same slice are stored sequentially in the disk.
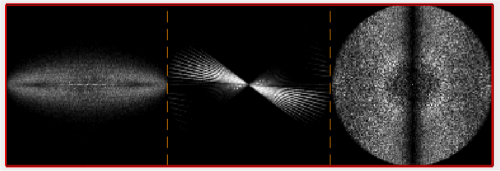
Estimation of the missing wedge
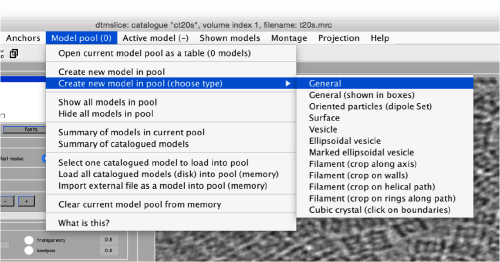
Creating a template
There are different strategies to create the first template. In the case where the general shape of each protein is roughly recognisable by eye, it is not difficult to just crop and align manually some of the particles. When this is not possible, you have the option of using a density map that mimics the general topology of your protein.
Through manual alignment
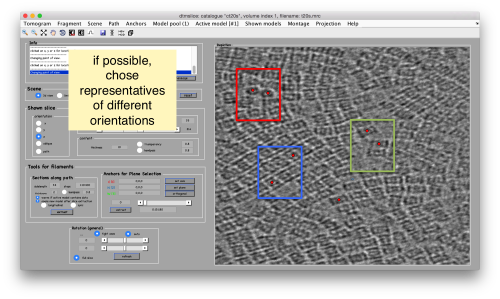
Manual selection of some particles
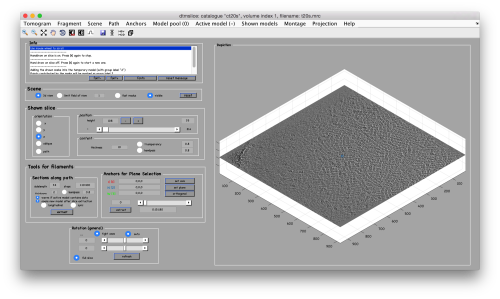
We use for this our tool dtmslice. As the tomogram is provided is fairly small you can probably just open it without any further binning.
dtmslice t20s.mrc -cc ct20
Manual alignment of some particles
[[ |thumb|center|500px ]]
[[ |thumb|center|500px ]]
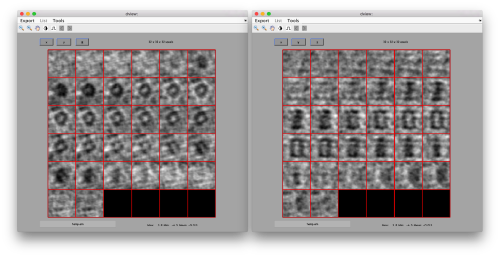
Cropping the borders of a template
Through geometrical shapes
Alternatively, you can use dynamo_mask or dynamo_tube to create a synthetic model.
Creating a cross correlation process
<notwiki>ptm = dynamo_match(fileFull,'average48.em','mask','maskData48.em',.... 'outputFolder','cs30',... 'ytilt',[-39,36],'sc',[1000,1000,300],'cr',360,'cs',30,'bin',1);<notwiki>
<notwiki>------------------------------------------------------------
Template matching process. computing in CPU Output folder: cs30.TM
A total of 1 tiles have been created
- Mb per tile (reading) : 724.19 - Mb per tile (operation) : 663.84 ... initializing output elements
Preparing to run on 1 blocks Running on single processor
Computing tile 1
-Range (original block): x[1:958] y[1:926] z[1:214]
-Range (binned block) : x[1:479] y[1:463] z[1:107]
... tile 1 finished in 387 seconds (8 for setup; 7.73 per triplet)
[ok] ... template matching process completed upto creation of cross correlation
you can proceed to peak location and particle extraction.
</notwiki>
The Process object
The return of the function <notwiki>ptm</nowiki> is an object of class XXX. In short, this is an object created
Considerations when creating a process
Locating cross correlation peaks
Looking at the cross correlation map
Looking at the cross correlation profile
Extracting a table
A table can be extracted through:
myTable = pff.peaks.computeTable('mcc',0.378);
Visual evaluation of results
Looking at cropped particles
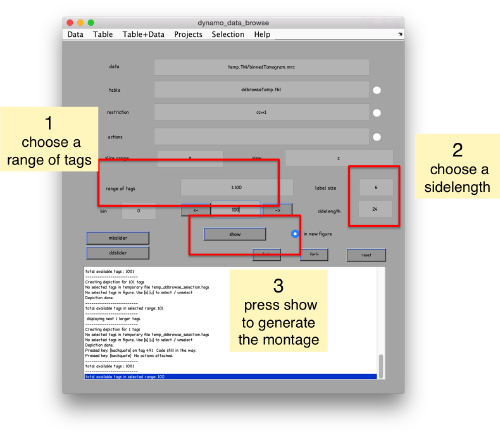
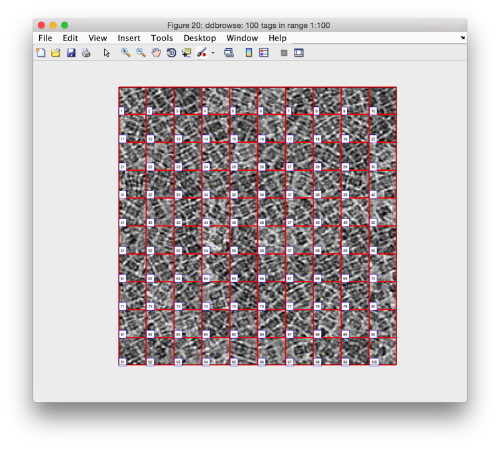
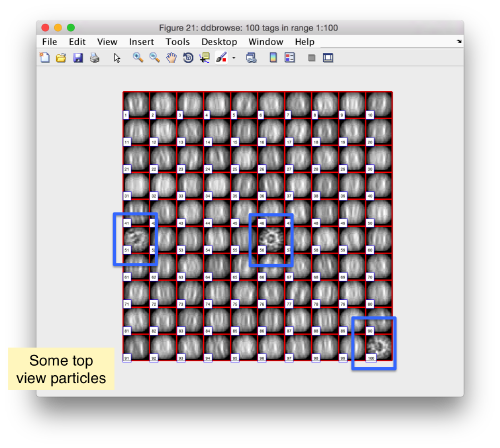
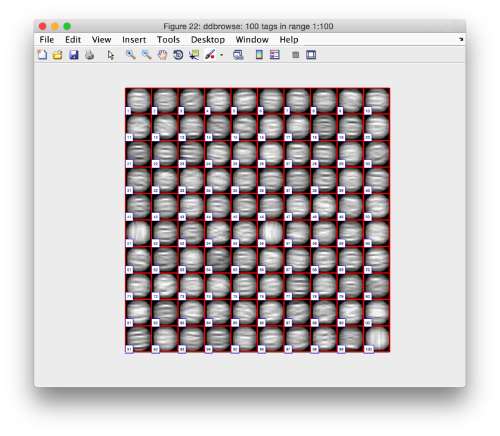
We can check how the individual particles look like on a gallery modus:
pff.peaks.browse();
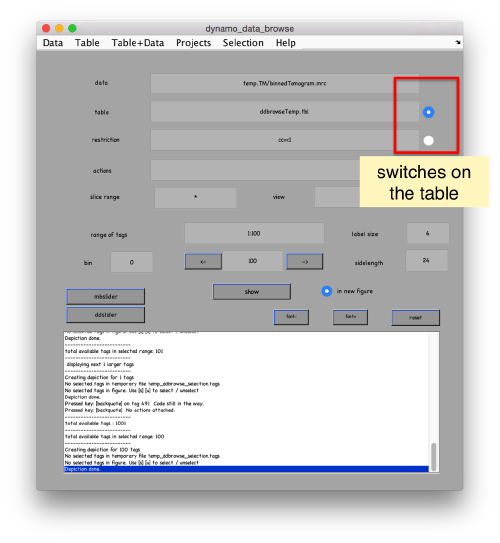
This order just opens ddbrowse. We are using here the support object peaks, but this command is equivalent to just invoking ddbrowse
ddbrowse('d','cr30.TM','t',myTable);
Looking at averages
o = pff.peaks.average();